A membrane-bound nuclease directly cleaves phage DNA during genome injection
TL;DR
SNIPE is a bacterial defense system that localizes to the cell membrane and directly cleaves phage DNA during genome injection, exploiting spatial organization to distinguish self from non-self. It associates with host proteins like ManYZ and phage tape measure proteins to target viral DNA specifically.
Key Takeaways
- •SNIPE is a membrane-bound nuclease in E. coli that provides direct defense by cleaving phage DNA during genome injection, preventing infection without harming host cells.
- •The system localizes to the inner membrane to avoid autoimmunity, with its transmembrane domain, DUF4041 domain for DNA binding, and GIY-YIG nuclease domain being essential for function.
- •SNIPE associates with host proteins ManYZ and phage tape measure proteins, enabling it to target diverse siphoviruses by exploiting the spatial organization of phage genome entry.
- •Defense is specific to phage infection, as SNIPE does not affect plasmid transformation or cleave host DNA when properly localized, representing a novel prokaryotic immune strategy.
Tags
Abstract
From mammals to bacteria, the direct recognition and cleavage of viral nucleic acids is a potent defence strategy against viral infection, but it requires mechanisms for distinguishing self from non-self1,2. In bacteria, CRISPR–Cas and restriction-modification systems achieve this discrimination by recognizing specific DNA sequences or DNA modifications, respectively. Alternative mechanisms probably remain to be discovered. Here, we characterize SNIPE, an anti-bacteriophage defence system that constitutively localizes to the bacterial cell membrane in Escherichia coli to block phage λ infection. Using radiolabelled phage DNA and time-lapse microscopy to track phage genomes, we demonstrate that SNIPE directly cleaves phage DNA during genome injection. Based on proximity labelling, we find that SNIPE associates with host proteins essential for λ genome entry and with the λ tape measure protein, which facilitates λ genome injection across the inner membrane. SNIPE also defends against diverse siphoviruses, probably through direct interactions with their tape measure proteins. Our findings establish SNIPE as a widespread bacterial defence system that exploits the spatial organization of phage genome injection to specifically target viral DNA, representing a previously unknown strategy for distinguishing self from non-self in prokaryotic immune systems.
Similar content being viewed by others
The coordination of anti-phage immunity mechanisms in bacterial cells
Continuous multiplexed phage genome editing using recombitrons
The ϕPA3 phage nucleus is enclosed by a self-assembling 2D crystalline lattice
Main
The ability to distinguish self from non-self is a fundamental feature of immune systems across all domains of life. In eukaryotes, pattern-recognition receptors enable this distinction by detecting conserved pathogen-associated features, such as lipopolysaccharides and flagellin3,4,5,6. Activated pattern-recognition receptors can trigger various immune responses, including the production of pro-inflammatory cytokines and the initiation of programmed cell death. Similarly, bacteria use abortive infection systems that recognize conserved features of invading bacteriophages and activate effectors that kill or arrest the host cell to prevent phage spread7,8,9,10.
A second form of non-self recognition involves targeting foreign nucleic acids directly. In eukaryotes, the RNA interference pathway cleaves viral double-stranded RNA into small interfering RNAs that then guide Argonaute proteins to complementary RNA sequences, leading to the degradation of viral RNA1,11. In bacteria, various ‘direct defence’ systems also specifically target foreign DNA. This includes CRISPR–Cas systems and Argonautes that use guide RNAs or DNAs, respectively, to cut foreign DNA in a sequence-specific manner1,2. Furthermore, restriction-modification systems can recognize foreign DNA based on the presence or absence of DNA modifications12. Whether there are other mechanisms for directly identifying and degrading foreign nucleic acids has remained an open question.
SNIPE is a membrane-bound nuclease
A previous genetic screen identified a direct defence system, provisionally named PD-λ-1, that potently blocks phage λ infection in E. coli13. For reasons described below, we renamed this system surface-associated nuclease inhibiting phage entry (SNIPE). To confirm that SNIPE provides direct defence, we infected cells at a concentration of phage λ such that approximately half of the cells were infected, and then monitored cell growth by time-lapse microscopy. In a population of cells lacking SNIPE, infected cells burst and caused a second round of successful phage infection in neighbouring cells, as expected (Fig. 1a and Supplementary Video 1). In sharp contrast to this, in a population of cells that did harbour SNIPE, no cell lysis was observed and cells grew to confluence. As a control, we also infected a population of cells containing an abortive infection system, PD-λ-3, and observed that infected cells lysed but prevented a second round of infection13. Moreover, growth-curve assays showed that SNIPE, but not PD-λ-3, enabled cell survival in the presence of high phage concentrations (Fig. 1b). These results confirm that SNIPE provides direct defence, enabling cells to ward off infection without compromising cell growth. Notably, SNIPE did not block plasmid DNA transformation, indicating that its activity is specific to phage infection (Extended Data Fig. 1a).
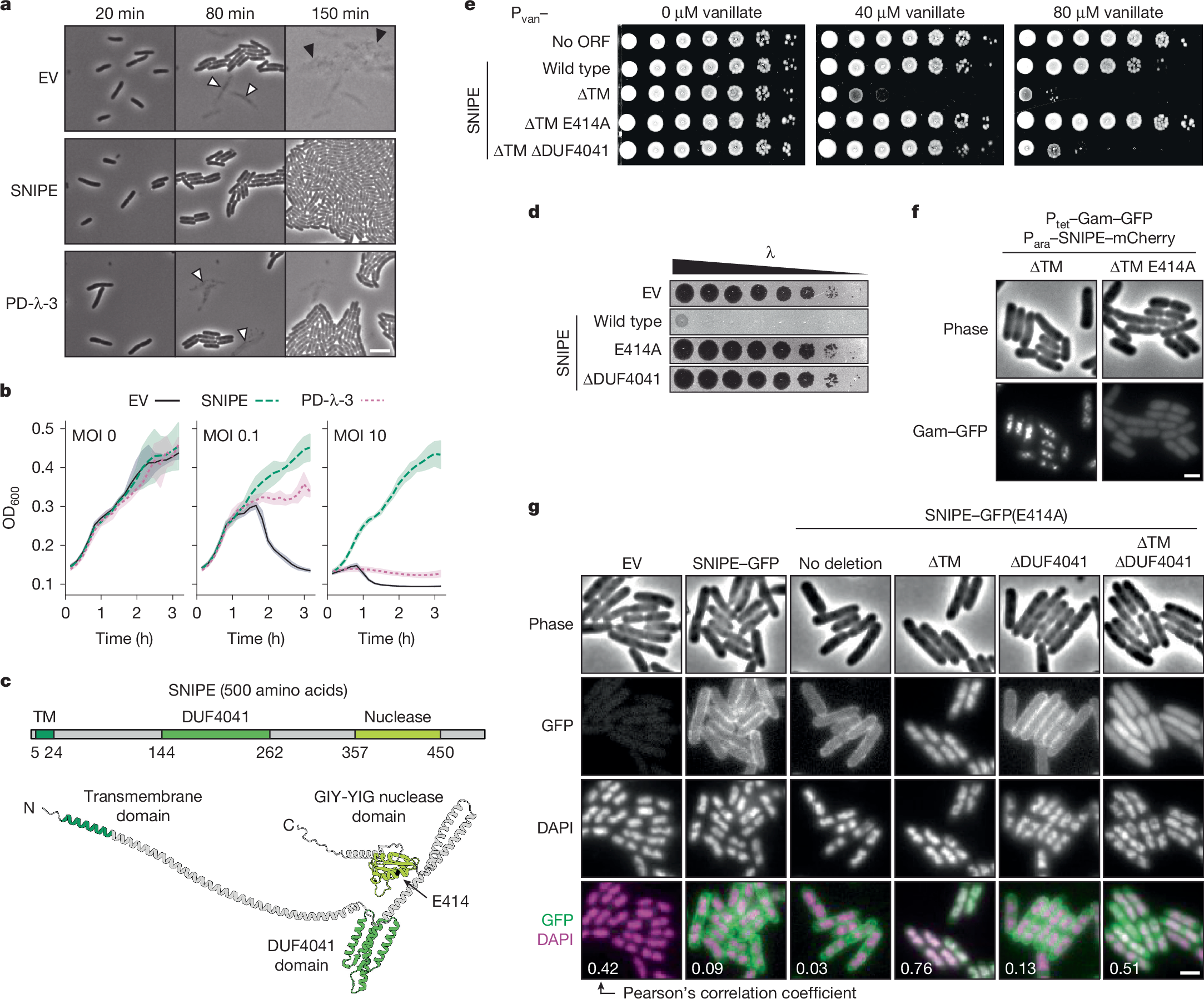
a, Time course of cells expressing an empty vector (EV), SNIPE or PD-λ-3. About half of the cells were infected with λ at time t = 0 min. White arrows show cell death from the initial round of infection and black arrows show cell death from the second round. Images are representative of n = 3 independent biological replicates. Scale bar, 3 µm. ORF, open reading frame. b, Growth curves for cells expressing an empty vector, SNIPE or PD-λ-3 and infected at different multiplicities of infection (MOI) of λ at t = 0 min. Optical density at 600 nm (OD600) was used as a proxy for bacterial cell density and growth. The line shows the mean and the shaded region is the standard deviation; n = 4 independent biological replicates. c, Structure of SNIPE predicted by AlphaFold3, with colour-coded domains predicted by HHpred and DeepTMHMM. The N terminus, C terminus and E414A location are indicated. The start and end points of each domain are shown. d, Serial dilutions of λ spotted on lawns of cells expressing an empty vector or different SNIPE constructs. e, Serial dilutions of bacterial strains on plates with 0, 40 or 80 µM vanillate to induce empty vector or Pvan–SNIPE constructs. f, Single-frame microscopy of cells expressing SNIPE–mCherry constructs and Gam–GFP, a marker of double-strand breaks. Images are representative of n = 3 independent biological replicates. Scale bar, 1 µm. g, Single-frame microscopy of cells expressing empty vector or SNIPE–GFP constructs. Nucleoids were stained with DAPI. Pearson’s correlation coefficients (R) were calculated for each condition by comparing the GFP and DAPI fluorescence signals (n > 100 cells). Scale bar, 1 µm.
Analyses using AlphaFold and HHpred indicated that SNIPE is an elongated protein with a transmembrane domain near the amino terminus, a domain of unknown function (DUF4041) in the middle of the protein, and a GIY-YIG nuclease domain near the carboxy terminus (Fig. 1c and Extended Data Fig. 1b,c). A transmembrane topology model predicted that the N terminus of SNIPE is periplasmic, the single-pass transmembrane domain resides in the inner membrane, and the rest of the protein is cytoplasmic14 (Extended Data Fig. 1d). We tested this model by inserting a PhoA–LacZα fusion protein at different locations in SNIPE15. PhoA functions only in the periplasm and LacZα functions only in the cytoplasm, so the respective activities of these domains indicate the topological orientation of SNIPE. PhoA was functional only when inserted at the N terminus of SNIPE, and LacZα was functional only when inserted downstream of the transmembrane domain (Extended Data Fig. 1e). These data indicate that SNIPE is anchored in the inner membrane, with the DUF4041 and nuclease domains protruding into the cytoplasm.
To test whether the transmembrane, DUF4041 and GIY-YIG nuclease domains are necessary for defence against phage λ, we mutated each of these regions individually and performed plaquing assays. Substituting a predicted catalytic residue in the nuclease domain (E414A) and deletion of the DUF4041 both abolished defence, indicating that these domains are essential for SNIPE function (Fig. 1d and Extended Data Fig. 1f). Attempts to clone SNIPE lacking the transmembrane domain (∆TM) were unsuccessful, so we put SNIPE(∆TM) under the control of a vanillate-inducible (Pvan) promoter. Induced expression of this construct was highly toxic, and this toxicity was abolished by the E414A mutation (the toxicity was also partly reduced by ∆DUF4041, which is explored below) (Fig. 1e). These data indicated that SNIPE(∆TM) might localize to and cleave host DNA. To test this model, we used Gam–GFP, a fluorescently tagged RecBCD inhibitor that localizes to double-strand breaks in vivo16. We first confirmed that this marker was functional by expressing the restriction enzyme EcoRI fused to mCherry, which localized to and cut host DNA, as shown by Gam–GFP foci (Extended Data Fig. 1g). Similarly, we found that SNIPE(∆TM)–mCherry localized to and cleaved host DNA (Fig. 1f). Consistent with the notion that the E414A mutation disrupts nuclease activity, no double-strand breaks were generated by SNIPE(∆TM E414A)–mCherry.
To assess the subcellular localization of SNIPE and SNIPE(∆TM) under the control of its native promoter, we first fused GFP to the C terminus of SNIPE and confirmed that this fusion protein retained robust defence against λ (Extended Data Fig. 2a). We found that SNIPE–GFP was uniformly localized to the cell membrane independent of phage infection (Fig. 1g). To visualize SNIPE(∆TM)–GFP without the toxic effects of nuclease activity, we used the catalytically inactive variant SNIPE(∆TM E414A)–GFP. This construct no longer localized to the cell membrane and instead associated with bacterial DNA, as judged by colocalization with the DAPI-stained nucleoid (Fig. 1g and Extended Data Fig. 2b). To confirm these results, we separated the cytoplasmic and membrane fractions of cell lysates and found by immunoblotting that SNIPE(E414A)–GFP was strongly enriched in the membrane fraction, whereas SNIPE(∆TM E414A)–GFP was strongly enriched in the cytoplasm (Extended Data Fig. 2c). These results further indicate that the toxicity of SNIPE(∆TM) stems from it localizing to and cleaving the host genome. By extension, these data indicate that wild-type SNIPE localizes to the inner membrane to help to sequester its nuclease activity and prevent autoimmunity.
Given that bacterial DNA probably contacts the cell membrane during processes such as chromosome replication and segregation17, and that SNIPE is not intrinsically toxic to cells, we proposed that membrane-localized SNIPE does not cleave membrane-localized host DNA. To test this, we ectopically localized host DNA to the cell membrane by fusing two transmembrane domains from MalF to GFP–Fis, a fluorescently tagged DNA-binding protein. Strong expression of MalF(TM1-2)–GFP–Fis localized DAPI-stained DNA to the cell membrane and was toxic even in the absence of SNIPE (Extended Data Fig. 2d). Across a range of MalF(TM1-2)–GFP–Fis expression levels with varying degrees of toxicity, the presence of SNIPE generated no extra toxicity for cells. This result indicates that localization of host DNA to the cell membrane does not subject it to SNIPE-mediated cleavage, indicating that membrane-localized SNIPE exists in an auto-inhibited state.
To understand why removing the DUF4041 reduced the toxicity of SNIPE(∆TM) (Fig. 1e), we examined the electrostatic surfaces of the predicted SNIPE structure and found that the DUF4041 domain contains a positively charged surface that might facilitate DNA binding (Extended Data Fig. 2e). To test this hypothesis, we compared the localization of SNIPE(∆TM E414A)–GFP with and without the DUF4041 domain. Indeed, whereas SNIPE(∆TM E414A)–GFP localized to the nucleoid, SNIPE(∆TM E414A ∆DUF4041)–GFP exhibited diffuse cytoplasmic localization (Fig. 1g). Furthermore, we found that a version of SNIPE containing only DUF4041 and the downstream α-helix fused to GFP localized to the bacterial nucleoid (Extended Data Fig. 2f). These observations indicate that the DUF4041 promotes DNA binding by SNIPE(∆TM). Collectively, our results support a model in which SNIPE contains a functional nuclease domain, a DUF4041 domain that facilitates DNA binding, and a transmembrane domain that anchors SNIPE to the inner membrane, preventing autoimmune cleavage of host DNA.
SNIPE cuts phage DNA during injection
Given that SNIPE is a membrane-bound nuclease, we reasoned that SNIPE could directly cleave phage DNA during genome injection. We first confirmed that SNIPE does not affect the adsorption of phage λ (Fig. 2a), consistent with a previous study13. Next, we sought to visualize genome injection directly using a fluorescence assay in which λ carried a parS site and E. coli produced CFP-ParB18. After phage-genome injection, CFP-ParB oligomerized on parS sites and λparS genomes appeared as fluorescent puncta. Consistent with previous work18, CFP-ParB puncta appeared within ten minutes of λparS infection and expanded during the course of phage-genome replication, which was followed by cell lysis (Fig. 2b,c and Supplementary Video 2). By contrast, the number of CFP-ParB puncta that appeared in SNIPE-expressing cells was reduced by about 30-fold, and there was a concomitant reduction in cell lysis. In rare cases in which a λparS genome appeared in a SNIPE-containing cell, it went on to replicate and lyse the cell, indicating that if phages stochastically evade SNIPE at the cell membrane, their development proceeds unimpeded (Extended Data Fig. 3a,b and Supplementary Video 2). The SNIPE-dependent reduction in CFP-ParB puncta was not observed with the catalytically inactive E414A mutation, indicating that this reduction was dependent on nuclease activity (Fig. 2b,c).
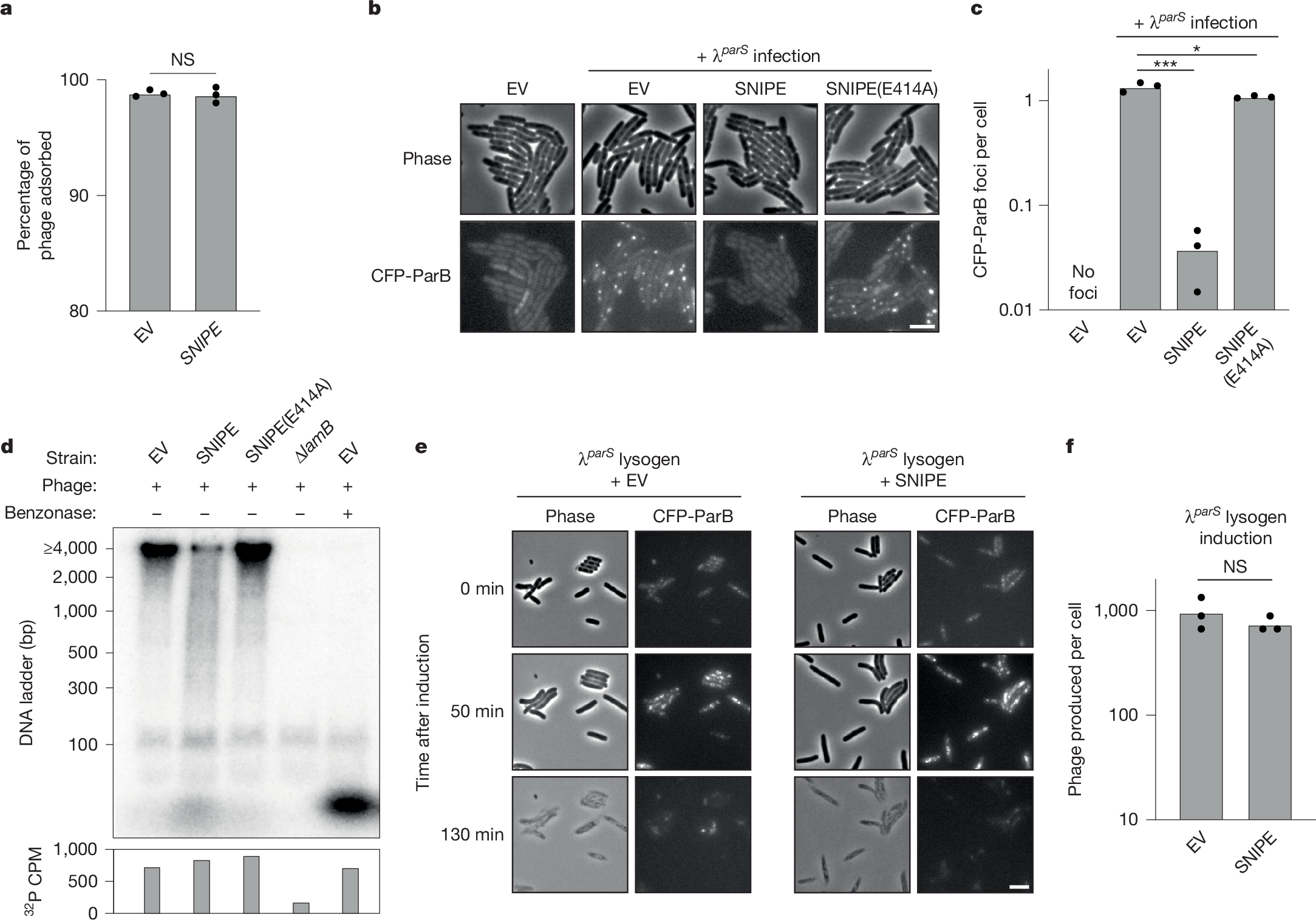
a, Adsorption assay with λ and cells containing SNIPE or an empty vector. Summary of n = 3 independent replicates. NS indicates P = 0.76 (unpaired two-tailed t-test). b, Microscopy of cells expressing CFP-ParB and an empty vector or different SNIPE constructs. Cells were infected with λparS and imaged 5 min after genome injection. Scale bar, 2 µm. c, Quantification of CFP-ParB foci per cell from b and two more independent replicates. *P = 0.03, ***P = 8.8 × 10−5 (unpaired two-tailed t-tests). d, Various bacterial strains were infected with 32P-labelled λ and lysed 15 min after genome injection. Benzonase was added to one lysate sample. Lysates were subjected to electrophoresis through a polyacrylamide gel and imaged with a phosphor screen. Total 32P per sample was measured with a scintillation counter. Data are shown from one biological replicate. Data from one other independent biological replicate are shown in Extended Data Fig. 3c. e, λparS lysogens expressing the heat-labile cI857 repressor, CFP-ParB and SNIPE or an empty vector were induced by heat shock at 42 °C. Time-lapse microscopy was used to monitor CFP-ParB foci dynamics and cell lysis. Scale bar, 2 µm. f, Quantification of plaque-forming units (PFU) per induced cell was done for the strains in e and two more independent replicates. NS, P = 0.35 (unpaired two-tailed t-test).
To test directly whether phage DNA is cleaved by SNIPE, we adapted the classic Hershey–Chase experiment19. In this adaptation, we infected cells with λ containing 32P-labelled DNA, collected cells shortly after genome injection and then measured the size of radiolabelled DNA fragments. Infection of empty vector cells yielded a clear 32P-labelled band at the upper limit of size detection (4,000 base pairs), as would be expected for the injected, roughly 42,000-bp λ genome (Fig. 2d). This band was cleaved into a band of less than 100 bp following benzonase treatment, and this smaller band probably represents mononucleotides because [γ−32P]-ATP formed a band of the same size (Extended Data Fig. 3c). Consistent with the 32P-labelled band originating from injected phage DNA, it was not observed following infection of cells lacking the λ receptor (∆lamB). In contrast to empty vector cells, infection of SNIPE-containing cells yielded a smear of DNA fragment sizes ranging from more than 4,000 bp to less than 100 bp, as well as the band that probably corresponded to mononucleotides. This profile of 32P-labelled DNA cleavage was reversed in cells expressing SNIPE(E414A). Importantly, the amount of injected 32P was similar across empty vector, SNIPE and SNIPE(E414A)-expressing cells, as measured by scintillation counting (Fig. 2d and Extended Data Fig. 3c). Taken together, our results strongly suggest that SNIPE cleaves phage DNA during the injection process.
We next investigated whether SNIPE could target a phage genome that was pre-existing in the cell. To this end, we introduced a plasmid expressing SNIPE or carrying an empty vector into a λparS lysogen that produces CFP-ParB and encodes a temperature-sensitive cI repressor, which causes the λparS prophage to enter the lytic cycle at high temperatures. Following heat shock, λparS prophages were induced and then replicated, as manifest by CFP-ParB foci, followed by synchronous cell lysis (Fig. 2e and Supplementary Video 3). As predicted, SNIPE did not qualitatively affect the dynamics of CFP-ParB foci or cell lysis, nor did it affect the number of phage particles produced by prophage induction (Fig. 2f). These findings support the model that SNIPE cleaves phage DNA during, but not after, genome injection.
SNIPE associates with ManYZ
We next wondered how SNIPE localizes to genome-injection sites. After adsorbing to the outer-membrane receptor LamB, phage λ requires the inner-membrane components of the mannose permease complex (consisting of ManY and ManZ) for genome injection, although the molecular details of this process remain poorly understood20,21. In principle, SNIPE could target phage DNA if it associates with LamB or ManYZ. After unsuccessful attempts to isolate λ escape mutants, we turned to a ‘generalist’ mutant of λ that evolved to infect ∆lamB and ∆manYZ strains22. This phage has a mutation in the tail tip protein that allows it to bind to either LamB or an alternative outer-membrane protein, OmpF (Fig. 3a). This phage also has a mutation in the tape measure protein that allows it to circumvent a requirement for ManYZ, although the nature of this alternative genome-injection pathway remains unclear.
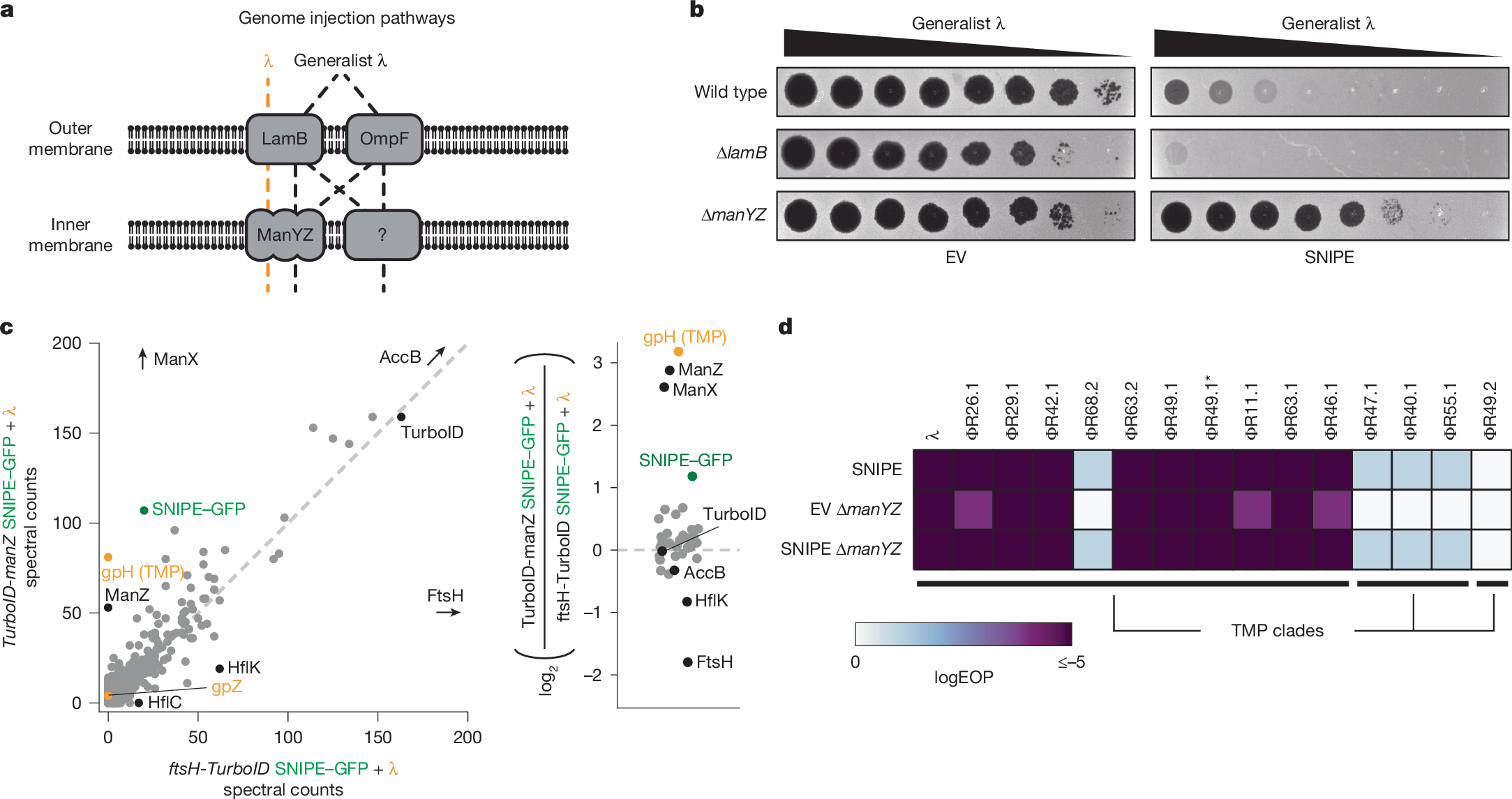
a, Schematic of genome injection routes used by λ and the generalist λ mutant. b, Serial dilutions of the generalist λ mutant were spotted onto bacterial lawns with different genomic deletions and an empty vector or SNIPE-expressing plasmid. c, Proximity labelling was done with infected cells expressing SNIPE–GFP and TurboID-ManZ or FtsH-TurboID. Biotinylated proteins were enriched with streptavidin beads and quantified by mass spectrometry. The log2 ratios of spectral counts for proteins with 50 or more spectral counts in at least one sample are shown on the right. A pseudocount was added to each spectral count value to facilitate ratio calculations. Phage proteins are labelled in orange, SNIPE–GFP is labelled in green, and other proteins of interest are labelled in black. d, Efficiency of plaquing (EOP) data for λ and other temperate phages on wild-type or ∆manYZ cells expressing SNIPE or harbouring an empty vector. Phages in a given TMP clade share more than 90% amino acid identity between their TMPs, and less than 20% identity with TMPs outside the clade.
We first confirmed that the generalist λ mutant was able to infect ∆lamB and ∆manYZ strains, albeit with minor plaquing defects (Fig. 3b). Next, we reasoned that if SNIPE-mediated defence requires LamB or ManYZ, then shunting genome injection through one of the alternative pathways would circumvent defence. SNIPE provided robust defence against the generalist λ mutant in both wild-type and ∆lamB cells, demonstrating that LamB is not required for defence. By contrast, SNIPE-mediated defence was strongly reduced in ∆manYZ. These data indicate that, although the generalist λ mutant can use various genome-injection pathways, ManYZ is the preferred pathway through the inner membrane if provided. As such, the generalist λ mutant probably uses