Convergent MurJ flippase inhibition by phage lysis proteins
TL;DR
Phage lysis proteins Sgl M, Sgl PP7, and Sgl CJ3 inhibit the bacterial cell wall target MurJ by trapping it in a periplasm-open conformation, revealing a convergent mechanism for antibiotic development.
Key Takeaways
- •MurJ is an essential bacterial lipid II flippase and a promising antimicrobial target.
- •Phage-encoded single-gene lysis proteins (Sgls) from M, PP7, and Changjiang3 phages inhibit MurJ despite no sequence similarity.
- •Structural studies show all three Sgls converge to trap MurJ in a periplasm-open conformation via a common interface.
- •This convergent inhibition mechanism provides a pathway for designing new antibiotics against drug-resistant bacteria.
Tags
Abstract
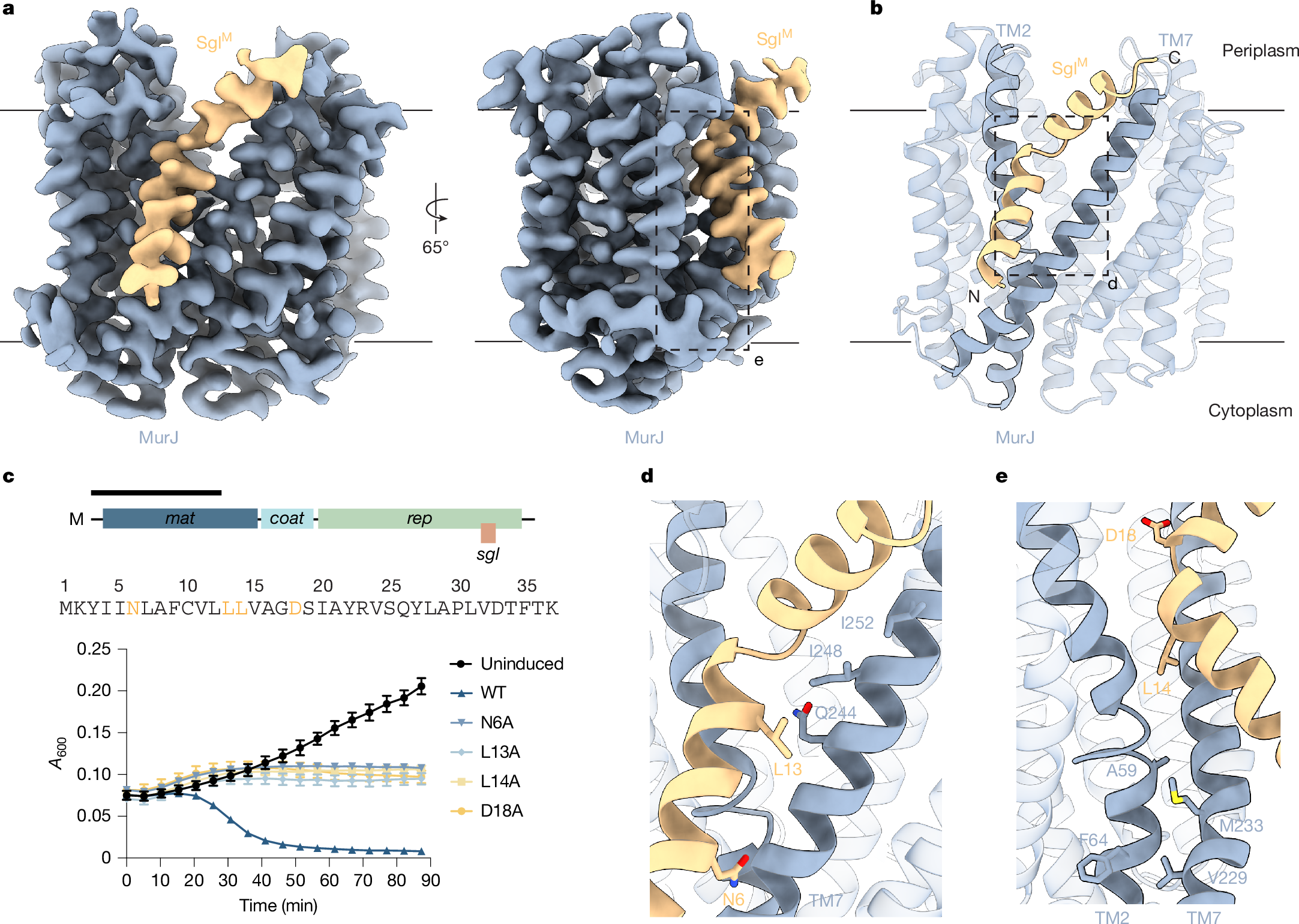
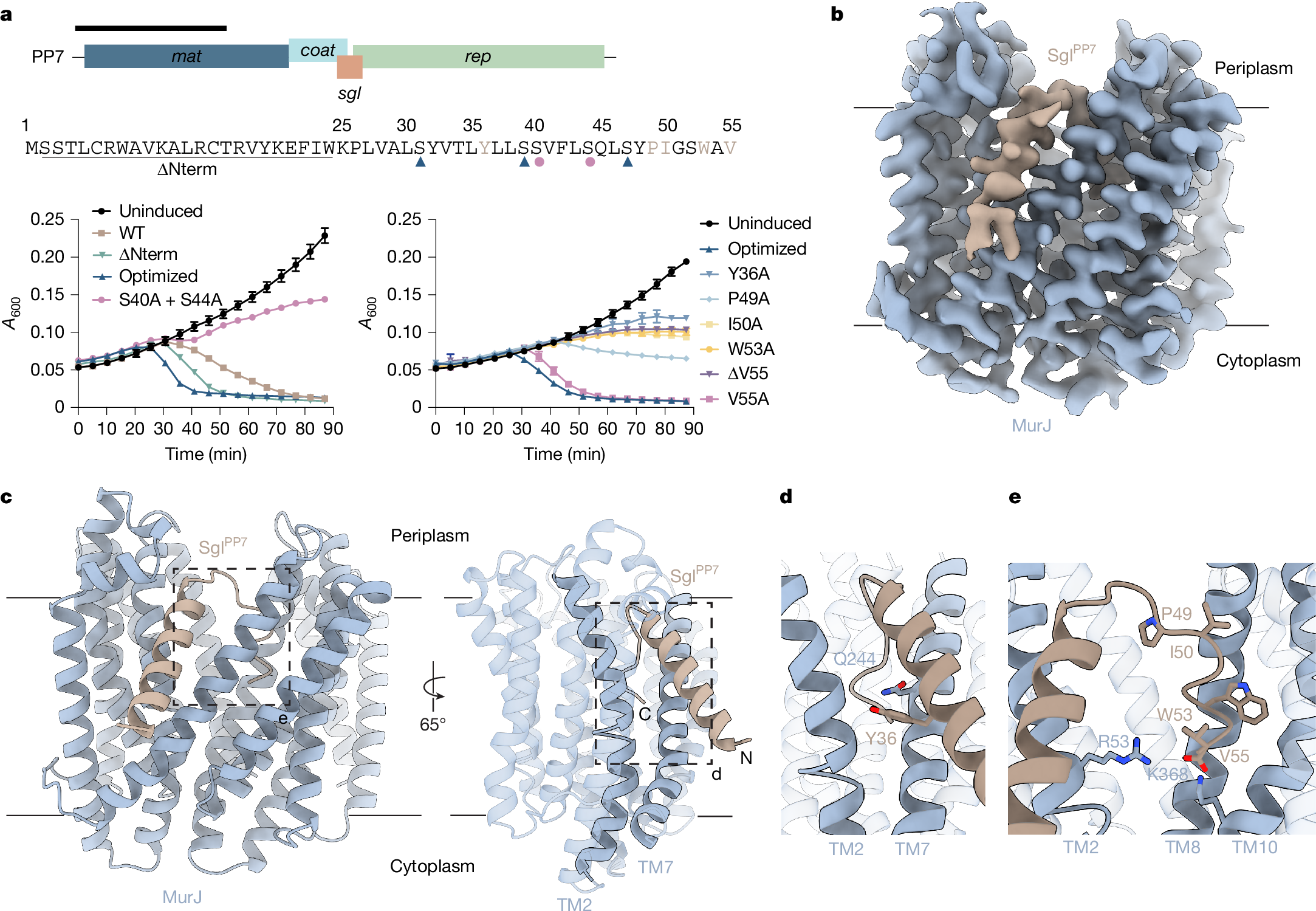
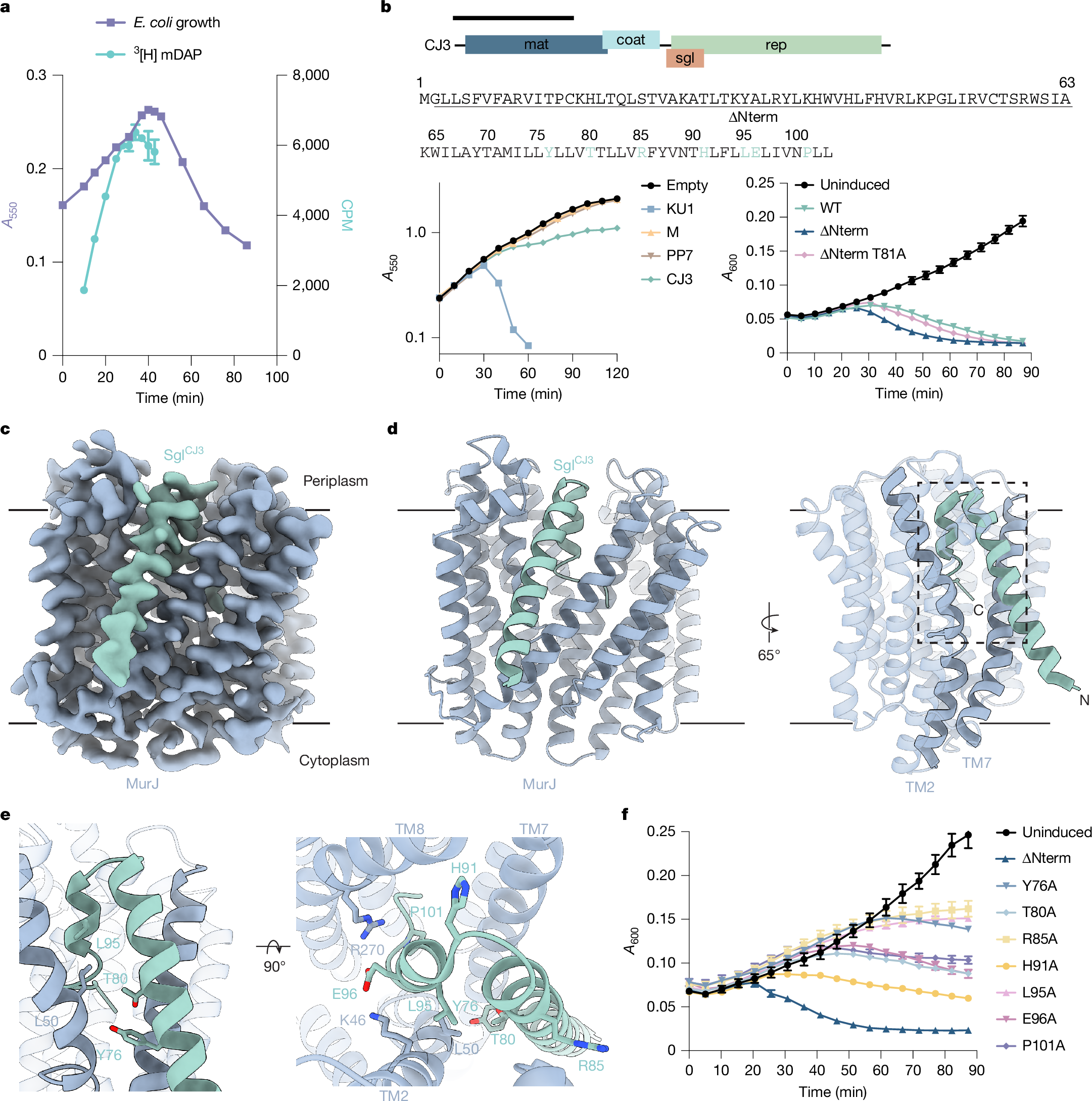
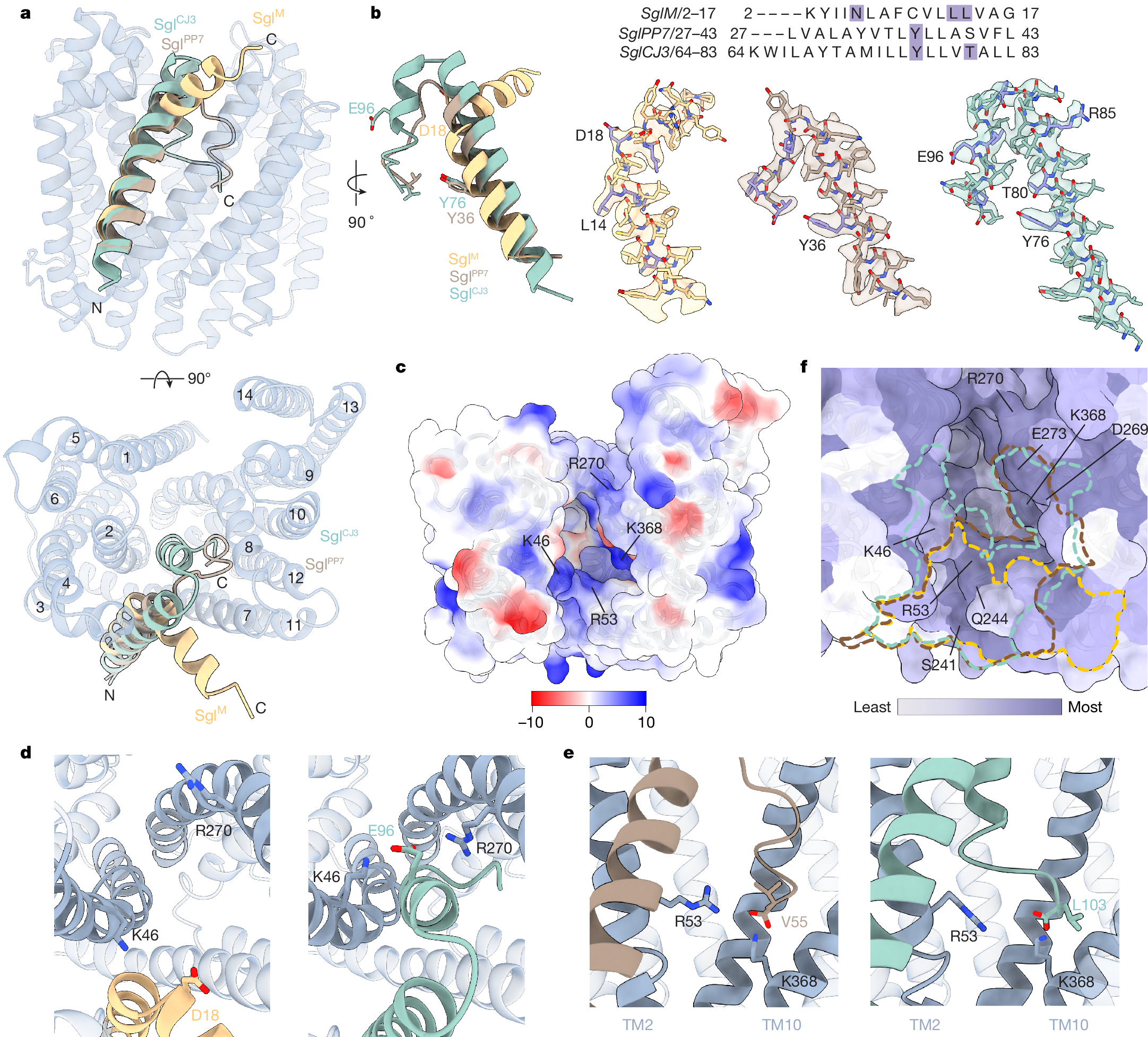
Antimicrobial drug resistance poses a global health challenge that necessitates the identification of new druggable targets1,2,3. The essential lipid II flippase MurJ is a promising yet underexplored antimicrobial target in bacterial cell wall biosynthesis4,5,6,7. The only known inhibitors of Gram-negative (diderm) MurJ are the single-gene lysis proteins (Sgls) from the lytic single-strand RNA phages M (SglM) and PP7 (SglPP7)8,9. SglM and SglPP7 have distinct evolutionary origins and share no sequence similarity. Here we describe a common mechanism of MurJ inhibition by these phage-encoded Sgls. We determined the structures of MurJ-bound SglM and SglPP7 and discovered a third distinct MurJ-targeting Sgl from the predicted phage Changjiang3 (SglCJ3) that we also characterized structurally. Our findings demonstrate that all three Sgls evolved convergently to trap MurJ in a periplasm-open conformation through a common MurJ interface, revealing a pathway for drug design.
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
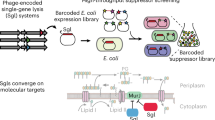
Multicopy suppressor screens reveal convergent evolution of single-gene lysis proteins

Antimicrobial potential of Streptomyces coeruleofuscus SCJ isolated from microbiologically unexplored garden soil in Northwest Morocco

Microbacterium kunmingensis sp. nov., an attached bacterium of Microcystis aeruginosa
Data availability
Atomic coordinates of SglM–MurJ, SglPP7–MurJ and SglCJ3–MurJ are deposited at the Protein Data Bank (PDB) with accession codes 9NU4, 9NU5 and 9NU8, respectively. Cryo-EM maps of SglM–MurJ, SglPP7–MurJ and SglCJ3–MurJ are deposited at the Electron Microscopy Data Bank (EMDB) with accession codes EMDB-49796, EMDB-49797 and EMDB-49798, respectively. Source data are provided with this paper.
Code availability
The code used for microscopy analysis of lysis morphology is available from GitHub at https://github.com/AntillonF/bleb_statistical_analysis.git.
References
Murray, C. J. L. et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399, 629–655 (2022).
Naghavi, M. et al. Global burden of bacterial antimicrobial resistance 1990–2021: a systematic analysis with forecasts to 2050. Lancet 404, 1199–1226 (2024).
Cook, M. A. & Wright, G. D. The past, present, and future of antibiotics. Sci. Transl. Med. 14, eabo7793 (2022).
Bugg, T. D. H., Braddick, D., Dowson, C. G. & Roper, D. I. Bacterial cell wall assembly: still an attractive antibacterial target. Trends Biotechnol. 29, 167–173 (2011).