Rewiring an E3 ligase enhances cold resilience and phosphate use in maize
TL;DR
Researchers identified the E3 ligase NLA as a key regulator linking cold stress and phosphate uptake in maize. By engineering a variant (NLA Δ12) that maintains cold tolerance while improving phosphate efficiency, they enhanced crop resilience and yield in field trials.
Key Takeaways
- •NLA E3 ligase regulates both cold tolerance (via JAZ11 degradation) and phosphate uptake (via PT4 degradation) in maize, creating a trade-off under cold stress.
- •A natural PT4 variant (K267A) was found to increase phosphate uptake by resisting NLA-mediated degradation in cold conditions.
- •AI-guided genome editing created the nla Δ12 allele, which uncouples these functions by impairing InsP binding while retaining JAZ11 targeting.
- •The engineered nla Δ12 maize shows improved cold resilience, higher phosphorus use efficiency (PUE), and increased grain yield in multi-site field trials.
- •This study provides a molecular framework for engineering climate-resilient, nutrient-efficient crops by tuning SPX regulatory modules.
Tags
Abstract
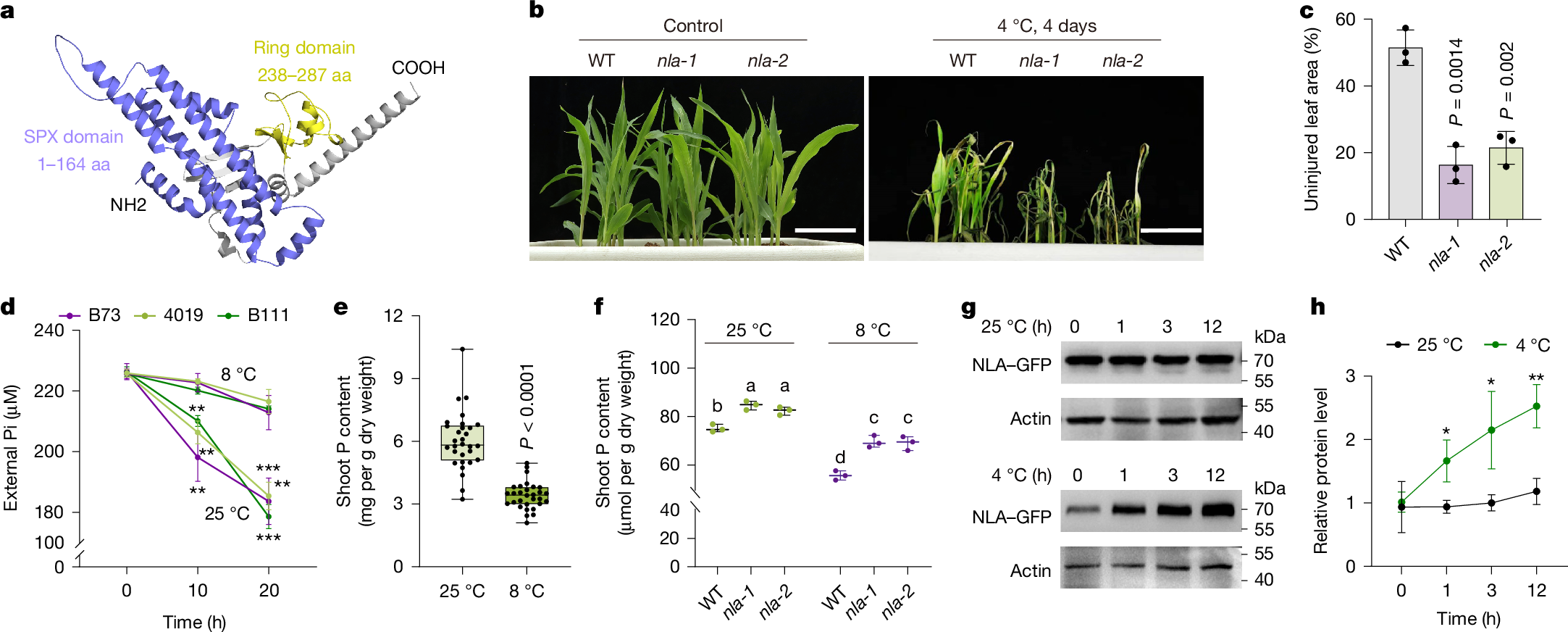
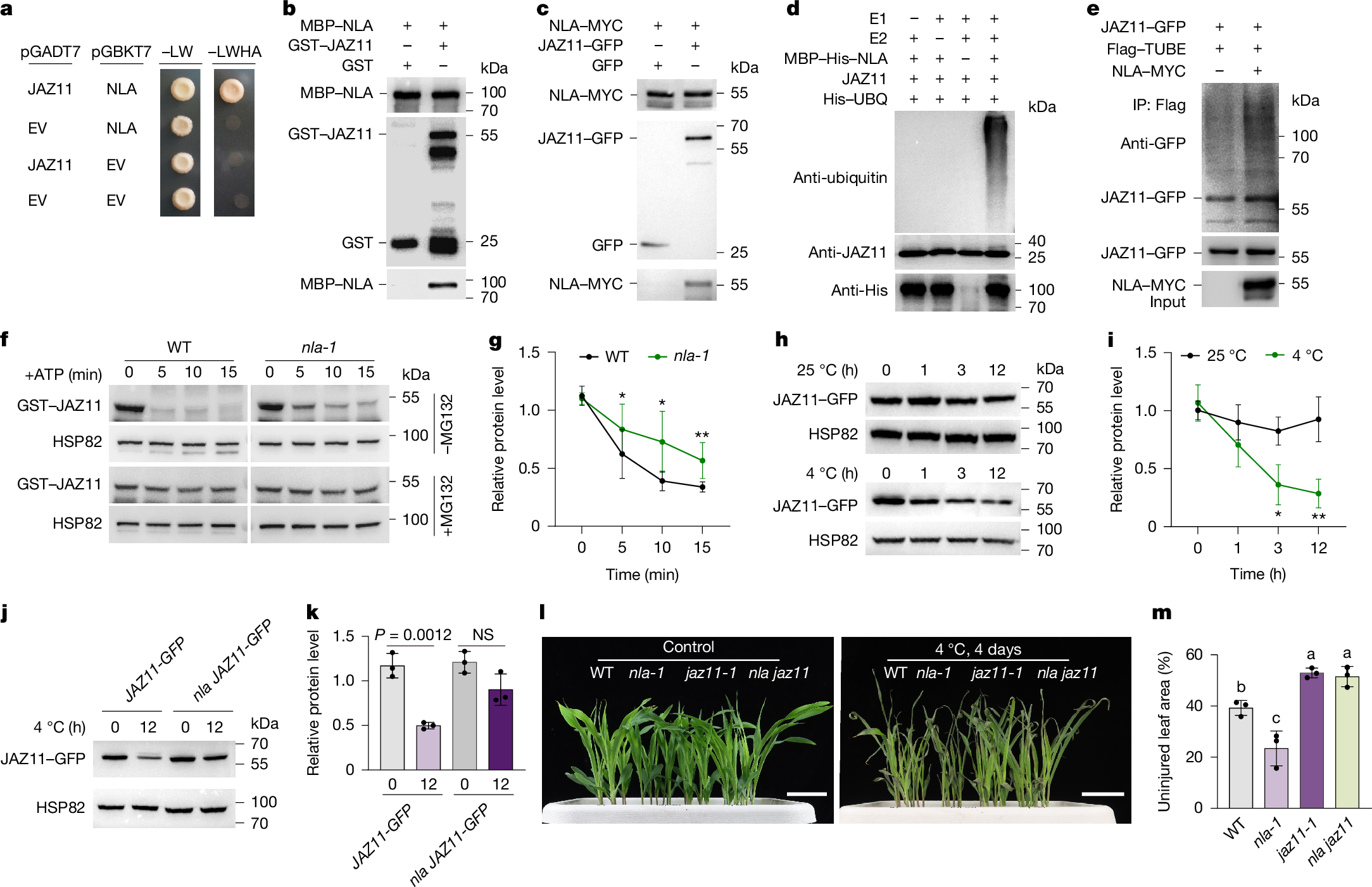
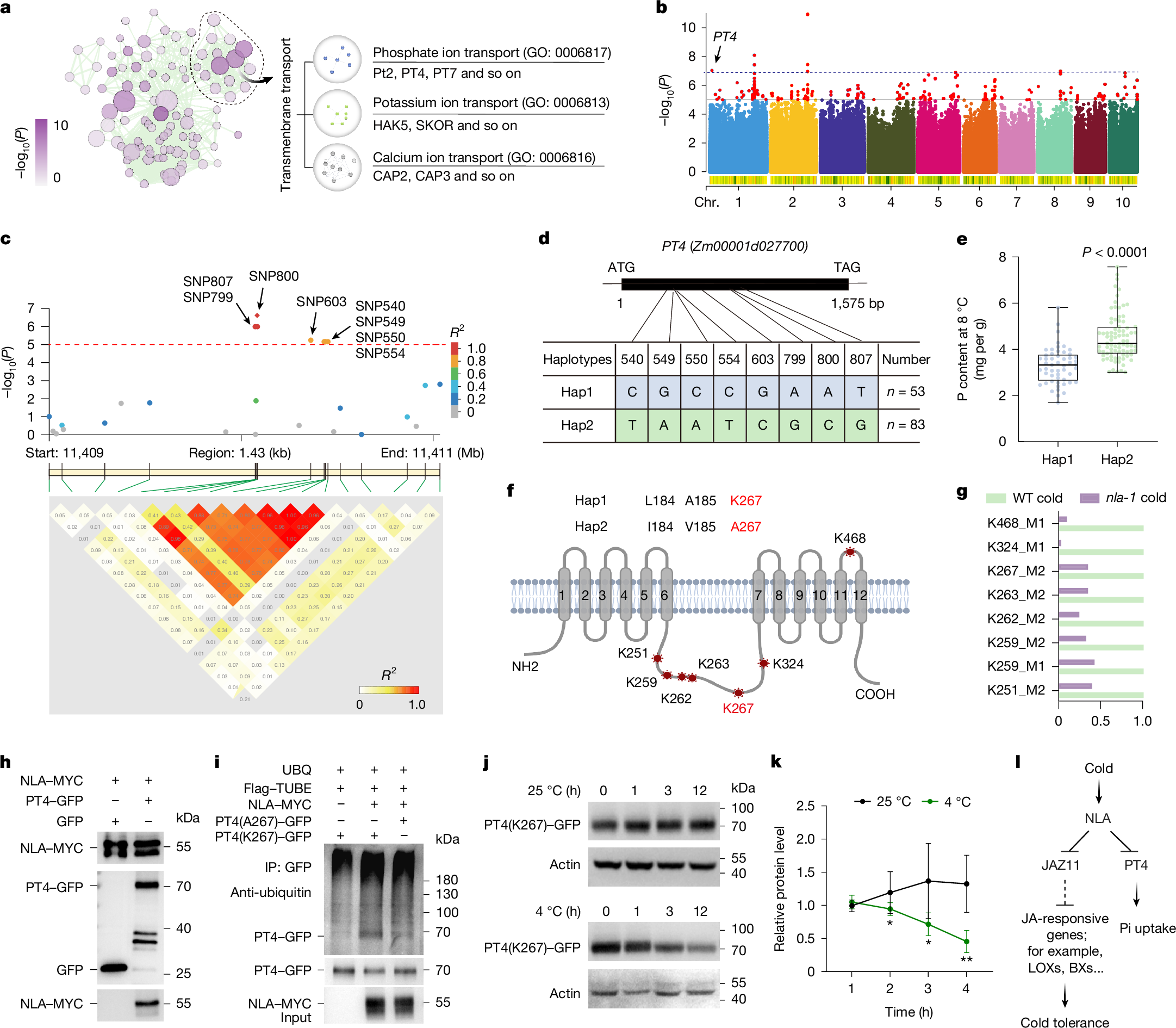
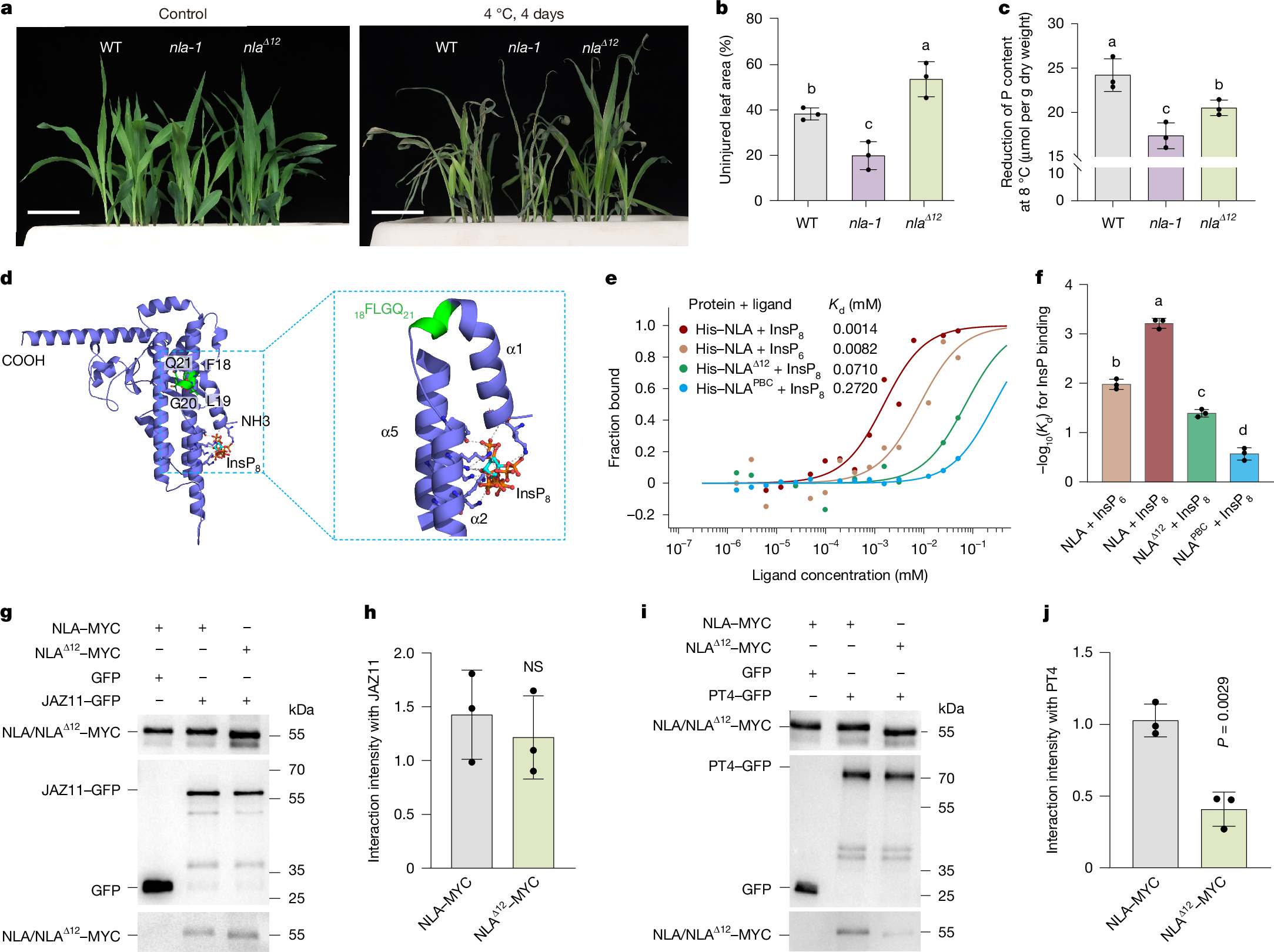
Cold stress restricts plant growth and inorganic phosphate (Pi) uptake, reducing yield and increasing fertilizer demand1,2,3. Enhancing both cold tolerance and phosphorus use efficiency (PUE) is crucial for sustainable crop productivity. Here we identify the SPX-domain-containing E3 ubiquitin ligase NITROGEN LIMITATION ADAPTATION (NLA) as a central regulator that links cold signalling to Pi homeostasis in maize (Zea mays L.). Under cold conditions, NLA promotes the degradation of the transcriptional repressor JAZ11, activating jasmonate signalling to enhance cold tolerance; however, NLA also simultaneously represses Pi uptake, through inositol polyphosphate (InsP)-dependent ubiquitination of the Pi transporter PT4. A ubiquitinome-informed genome-wide association study identified a natural PT4(K267A) (lysine-to-alanine substitution) variant that attenuates NLA-mediated degradation and increases Pi uptake in cold conditions. To overcome this nutrient–stress trade-off, we combined artificial-intelligence-guided structural modelling and ligand docking with genome editing to generate the nlaΔ12 allele, which encodes an NLA variant in which binding to InsP is impaired but JAZ11 targeting is retained. The Δ12 modification selectively redirects the activity of NLA towards jasmonate signalling, resulting in improved cold resilience, higher PUE and increased yield in multi-site field trials. These findings reveal a tunable SPX regulatory module that integrates environmental and nutrient signals, and provide a molecular framework for engineering climate-resilient, nutrient-efficient crops.
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others

ZmICE1a regulates the defence–storage trade-off in maize endosperm
Responses of rice plant to multiple abiotic stresses revealed by transcriptome meta-analysis and identification of novel genetic factors

Nature variations of OsNLP4 responsible for nitrogen use efficiency divergence in the two rice subspecies
Data availability
Gene sequences and amino acid sequences used in this study were collected from MaizeGDB (maize; https://www.maizegdb.org/), Gramene (rice; http://ensembl.gramene.org/index.html) and TAIR10 (Arabidopsis; https://www.arabidopsis.org/). The co-expression data used in this study are available from ATTED-II (https://atted.jp/). The maize and teosinte genotype data were sourced from the HapMap3 resource74, which is publicly accessible through the Panzea database (www.panzea.org). The geographical distribution map was generated using the R packages maps v.3.4.0 and ggplot2 v.3.4.2. MS-based proteomics and ubiquitinomics datasets have been deposited in the Integrated Proteome Resources (iProX) database under accession code IPX0011717000. RNA-sequencing data generated in this paper are accessible through the National Genomics Data Center (NGDC) under the BioProject accession number PRJCA041350 and raw data accession number CRA026553. Uncropped gel and blot images are provided in Supplementary Fig. 1. Source data are provided with this paper.
References
Bravo-F, P. & Uribe, E. G. Temperature dependence of the concentration kinetics of absorption of phosphate and potassium in corn roots. Plant Physiol. 67, 815–819 (1981).
Lambers, H. Phosphorus acquisition and utilization in plants. Annu. Rev. Plant Biol. 20, 17–42 (2022).
Yang, Z. et al. Genetic and molecular exploration of maize environmental stress resilience: toward sustainable agriculture. Mol. Plant 16, 1496–1517 (2023).
Doebley, J. The genetics of maize evolution. Annu. Rev. Genet. 38, 37–59 (2004).