Merlin: a computed tomography vision–language foundation model and dataset
TL;DR
Merlin is a 3D vision-language foundation model for abdominal CT scans that integrates volumetric imaging, EHR data, and radiology reports. It outperforms existing models across multiple tasks and demonstrates strong generalization in clinical settings.
Key Takeaways
- •Merlin addresses limitations of 2D VLMs by using 3D CT scans and comprehensive data without manual annotations.
- •It was trained on a large dataset of over 6 million images and evaluated on 752 tasks, showing high performance in diagnostic and prognostic applications.
- •The model and dataset are publicly released to support automated medical image analysis and reduce radiologist workload.
Tags
Abstract
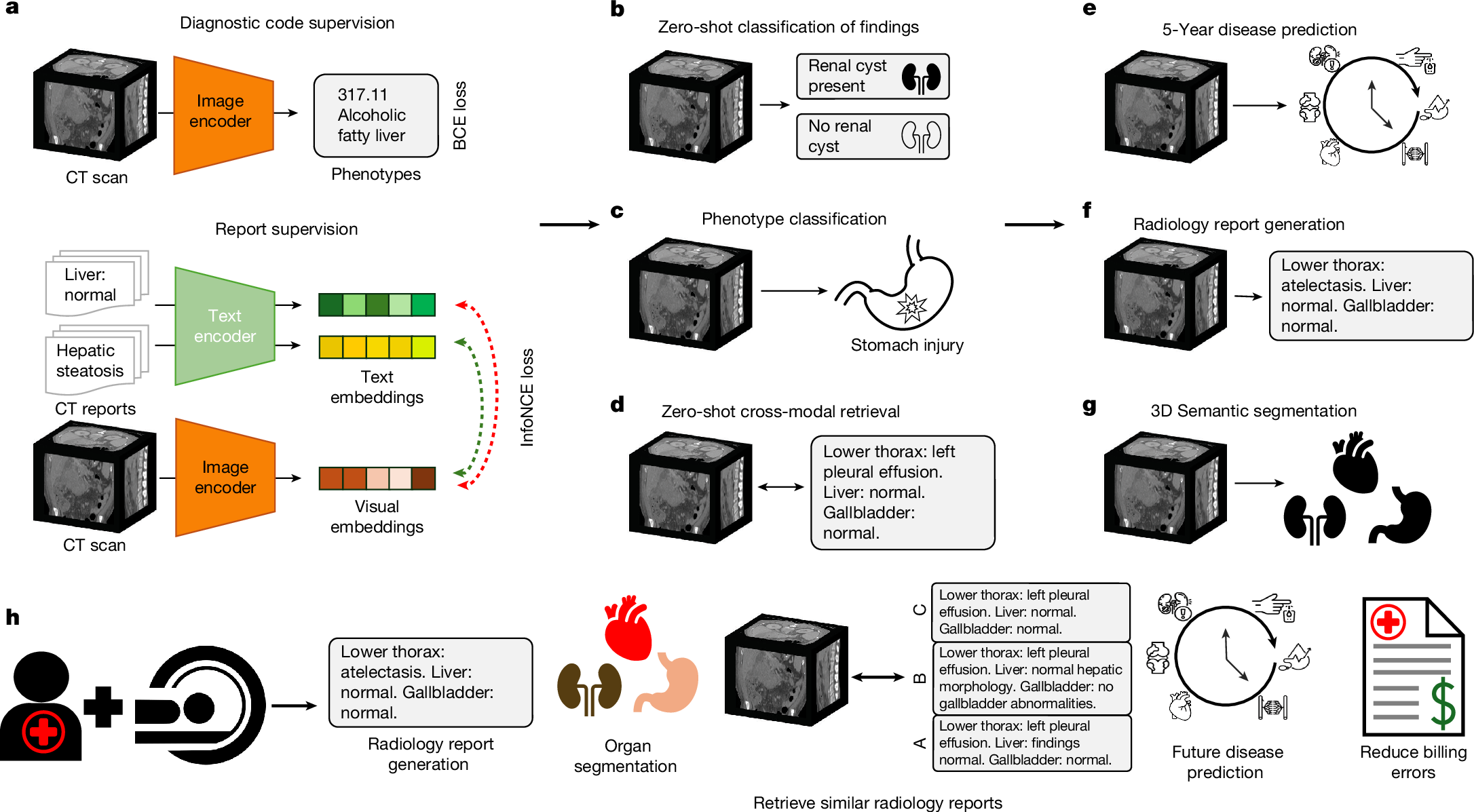
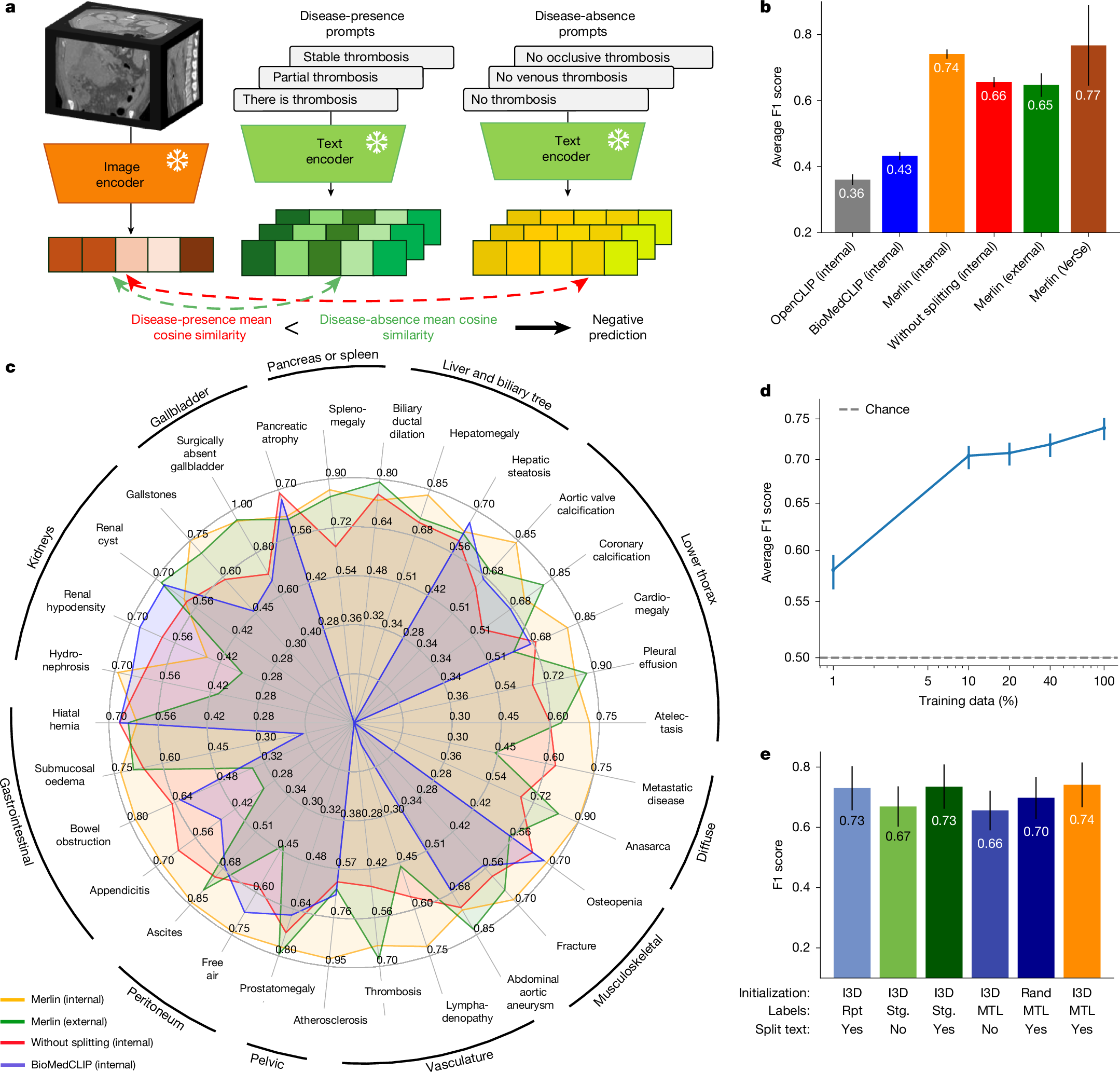
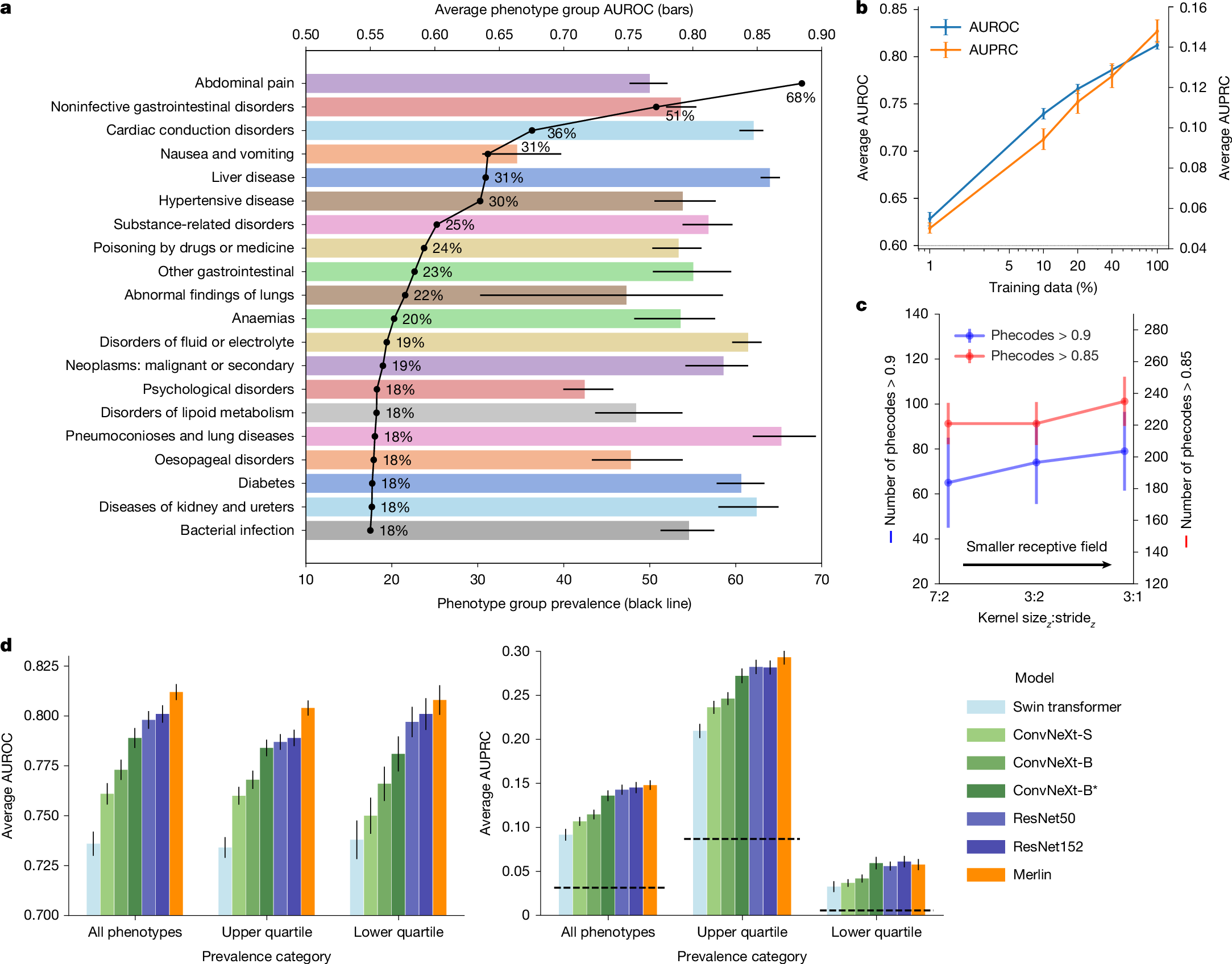
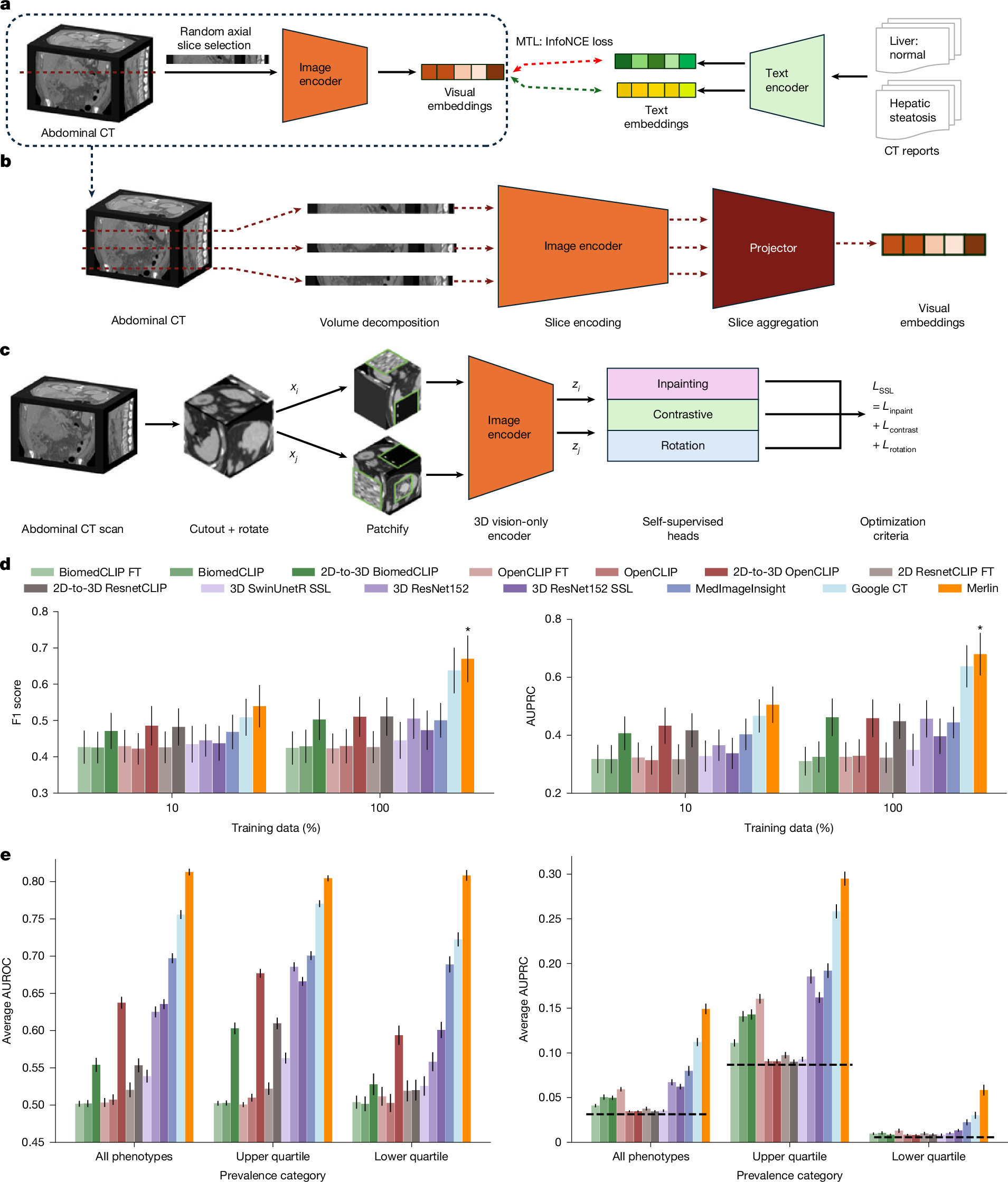
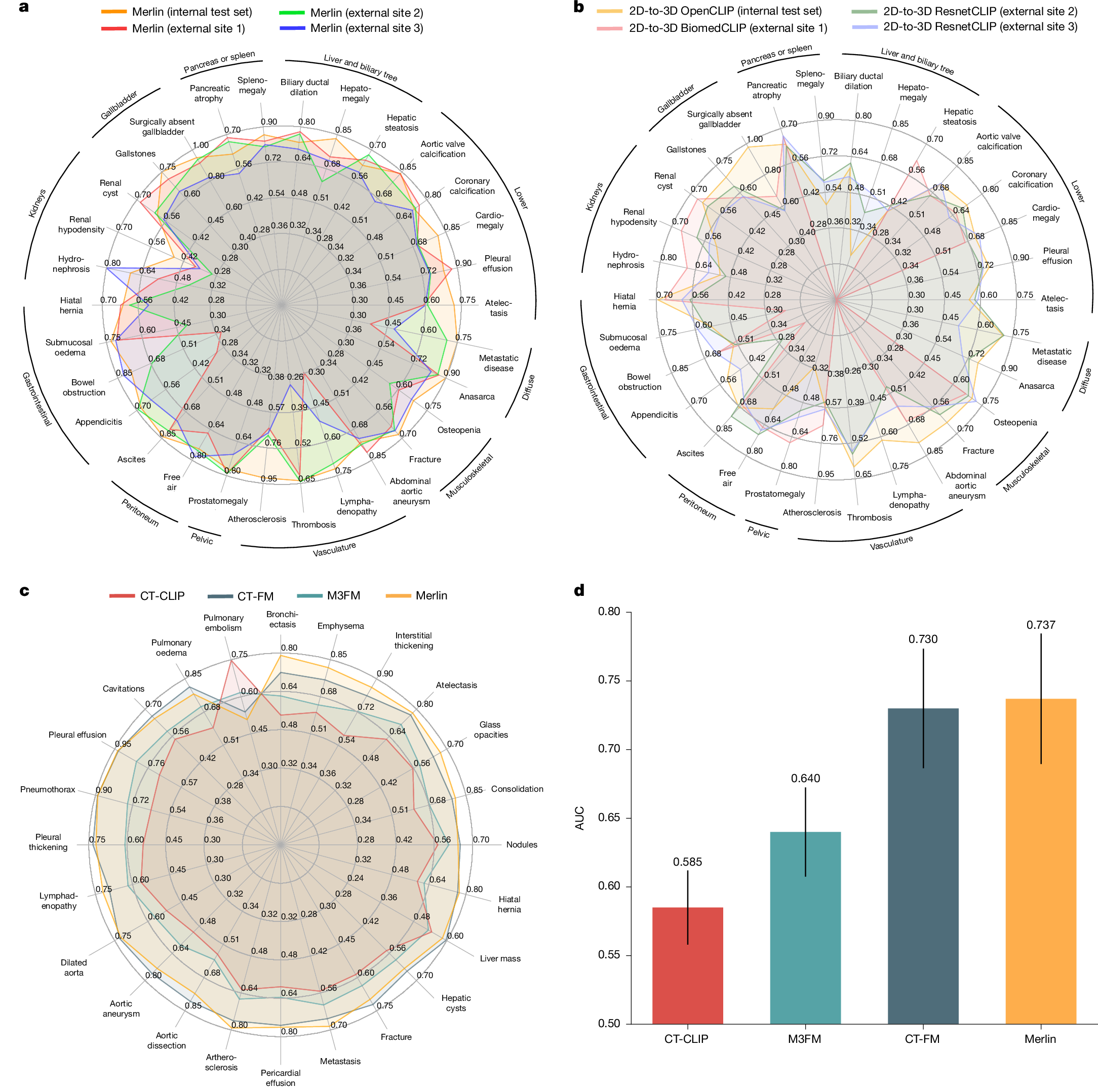
The large volume of abdominal computed tomography (CT) scans1,2 coupled with the shortage of radiologists3,4,5,6 have intensified the need for automated medical image analysis tools. Previous state-of-the-art approaches for automated analysis leverage vision–language models (VLMs) that jointly model images and radiology reports7,8,9,10,11,12. However, current medical VLMs are generally limited to 2D images and short reports. Here to overcome these shortcomings for abdominal CT interpretation, we introduce Merlin, a 3D VLM that learns from volumetric CT scans, electronic health record data and radiology reports. This approach is enabled by a multistage pretraining framework that does not require additional manual annotations. We trained Merlin using a high-quality clinical dataset of paired CT scans (>6 million images from 15,331 CT scans), diagnosis codes (>1.8 million codes) and radiology reports (>6 million tokens). We comprehensively evaluated Merlin on 6 task types and 752 individual tasks that covered diagnostic, prognostic and quality-related tasks. The non-adapted (off-the-shelf) tasks included zero-shot classification of findings (30 findings), phenotype classification (692 phenotypes) and zero-shot cross-modal retrieval (image-to-findings and image-to-impression). The model-adapted tasks included 5-year chronic disease prediction (6 diseases), radiology report generation and 3D semantic segmentation (20 organs). We validated Merlin at scale, with internal testing on 5,137 CT scans and external testing on 44,098 CT scans from 3 independent sites and 2 public datasets. The results demonstrated high generalization across institutions and anatomies. Merlin outperformed 2D VLMs, CT foundation models and off-the-shelf radiology models. We also computed scaling laws and conducted ablation studies to identify optimal training strategies. We release our trained models, code and dataset for 25,494 pairs of abdominal CT scans and radiology reports. Our results demonstrate how Merlin may assist in the interpretation of abdominal CT scans and mitigate the burden on radiologists while simultaneously adding value for future biomarker discovery and disease risk stratification.
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Data availability
We have released the Merlin abdominal CT dataset to the community (https://stanfordaimi.azurewebsites.net/datasets/60b9c7ff-877b-48ce-96c3-0194c8205c40). This large-scale abdominal CT dataset contains 25,494 scans of 18,317 unique patients, with each scan paired with its corresponding radiology report. Exams include abdominal and pelvis CT scans, identified using CPT codes 72192, 72193, 72194, 74150, 74160, 74170, 74176, 74177 and 74178, selected via the STARR tool (Stanford Medicine Research Data Repository). For each exam, the DICOM series with the largest slice count was retained and converted to NIfTI format for ease of use. Scans were compressed and de-identified by removing all patient-identifiable metadata. The Merlin abdominal CT dataset is hosted by the Stanford AIMI Center. Access requires completion of a data-use agreement form on the download page of the dataset. Following approval, a secure Azure Blob Storage URL is provided for download from the Merlin abdominal CT dataset download page. Additional download instructions are available in the documentation (download documentation). The following external publicly available datasets were used: the VerSe dataset is available via the Open Science Framework (https://osf.io/nqjyw/) and the TotalSegmentator dataset is publicly available at GitHub (https://github.com/wasserth/TotalSegmentator). Furthermore, the Merlin model was evaluated on abdominal CT images and associated radiology reports from three external clinical sites. These datasets are not publicly available owing to patient privacy considerations and data-use agreements but were accessed under appropriate institutional approvals and used solely for evaluation. All datasets were accessed and used in accordance with their respective data-use agreements and licences.
Code availability
Merlin is publicly available through the following platforms: GitHub (https://github.com/StanfordMIMI/Merlin), HuggingFace (https://huggingface.co/stanfordmimi/Merlin), and PyPI (https://pypi.org/project/merlin-vlm). The implementation builds on open-source libraries, including PyTorch (v.2.1.2), OpenCLIP (v.2.24.0) and the HuggingFace Transformers library (v.4.38.2). Model training was performed using the AdamW optimizer as implemented in PyTorch. For baseline comparisons, we used the BiomedCLIP model (hf-hub:microsoft/BiomedCLIP-PubMedBERT_256-vit_base_patch16_224) and OpenCLIP (hf-hub:laion/CLIP-ViT-L-14-laion2B-s32B-b82K) via the HuggingFace hub. Clinical text encoding was performed using the Clinical Longformer model (Yikuan8/Clinical-Longformer). The 3D inflation strategy for convolutional weights was adapted from the open-source repository at GitHub (https://github.com/hassony2/inflated_convnets_pytorch). The primary dataset used in this study is publicly available online (https://stanfordaimi.azurewebsites.net/datasets/60b9c7ff-877b-48ce-96c3-0194c8205c40).
References
Schöckel, L. et al. Developments in X-ray contrast media and the potential impact on computed tomography. Invest. Radiol. 55, 592–597 (2020).
Kanal, K. M. et al. U.S. diagnostic reference levels and achievable doses for 10 adult CT examinations. Radiology 284, 120–133 (2017).
Taschetta-Millane, M. The evolving computed tomography market. Imaging Technology News https://www.itnonline.com/article/evolving-computed-tomography-market (2024).
Hudnall, C. Maximum capacity: overloaded radiologists are grappling with solutions to a booming volume crisis. American College of Radiology https://www.acr.org/Practice-Management-Quality-Informatics/ACR-Bulletin/Articles/April-2024/Maximum-Capacity (2024).
Milburn, J. Workforce-shortage. How will we solve our radiology workforce shortage? American College of Radiology https://www.acr.org/Practice-Management-Quality-Informatics/ACR-Bulletin/Articles/March-2024/How-Will-We-Solve-Our-Radiology-Workforce-Shortage (2024).
Rimmer, A. Radiologist shortage leaves patient care at risk, warns royal college. BMJ 359, j4683 (2017).