A glucocorticoid–FAS axis controls immune evasion during metastatic seeding
TL;DR
A glucocorticoid receptor (GR) activation in disseminated tumor cells (DTCs) represses the FAS-FASL pathway, enabling immune evasion during metastatic seeding. GR inhibition combined with immunotherapy reduces metastasis in mice, suggesting a therapeutic target for early-stage metastatic cancer.
Key Takeaways
- •Glucocorticoid receptor (GR) activation in disseminated tumor cells (DTCs) drives resistance to CD8+ T cells and natural killer cells during early metastatic seeding.
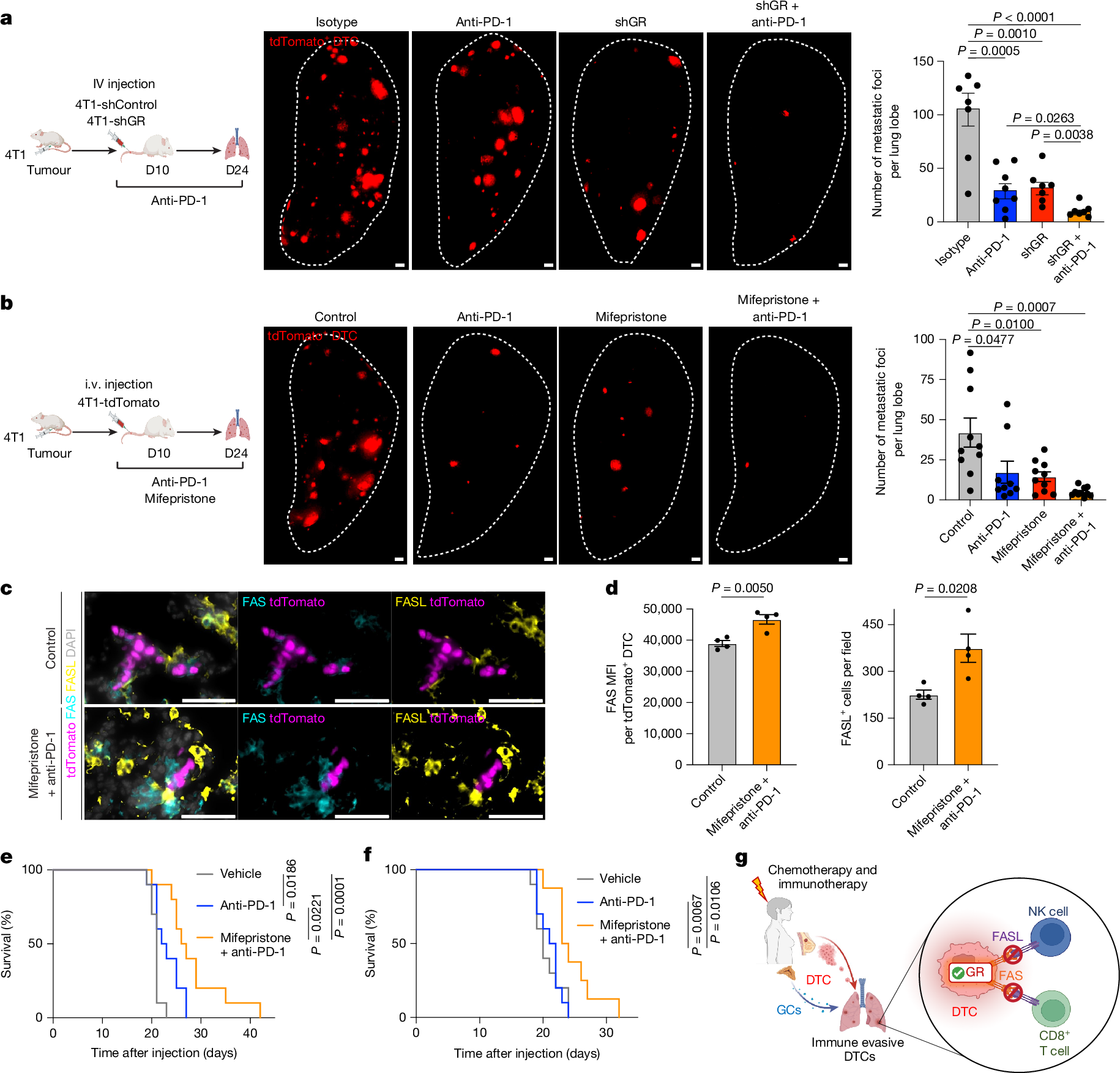
- •GR represses the FAS-FASL cytotoxic pathway, which is crucial for immune-mediated killing of DTCs in the metastatic niche.
- •Pharmacological inhibition of GR, when combined with immunotherapy (e.g., anti-PD-1), reduces metastatic burden and extends lifespan in mouse models.
- •This mechanism highlights unique immune-cancer interactions in DTCs, offering therapeutic opportunities distinct from primary tumor treatments.
- •The findings suggest GR inhibition as a promising target to eliminate DTCs and prevent metastasis in cancers like triple-negative breast cancer.
Tags
Abstract
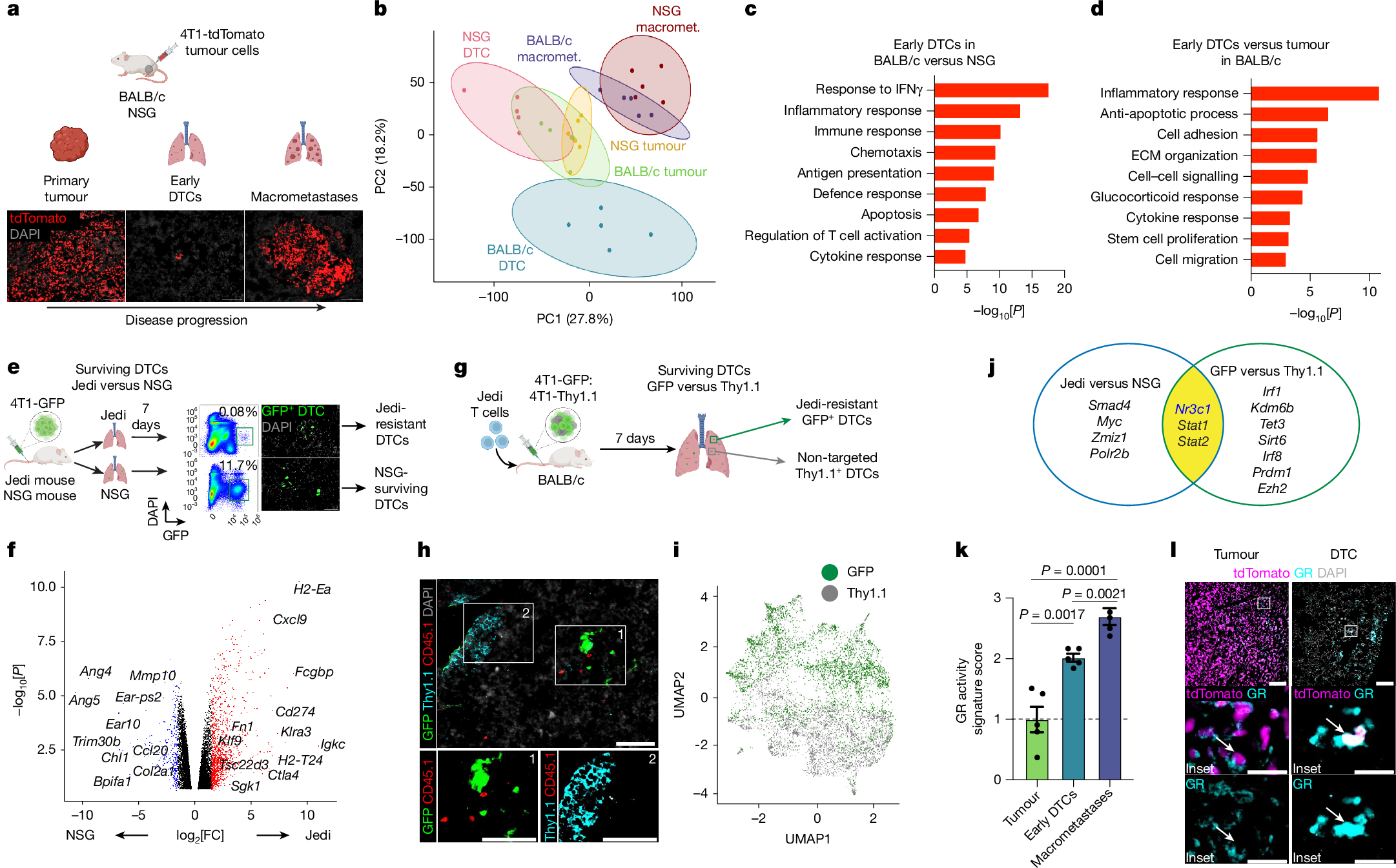
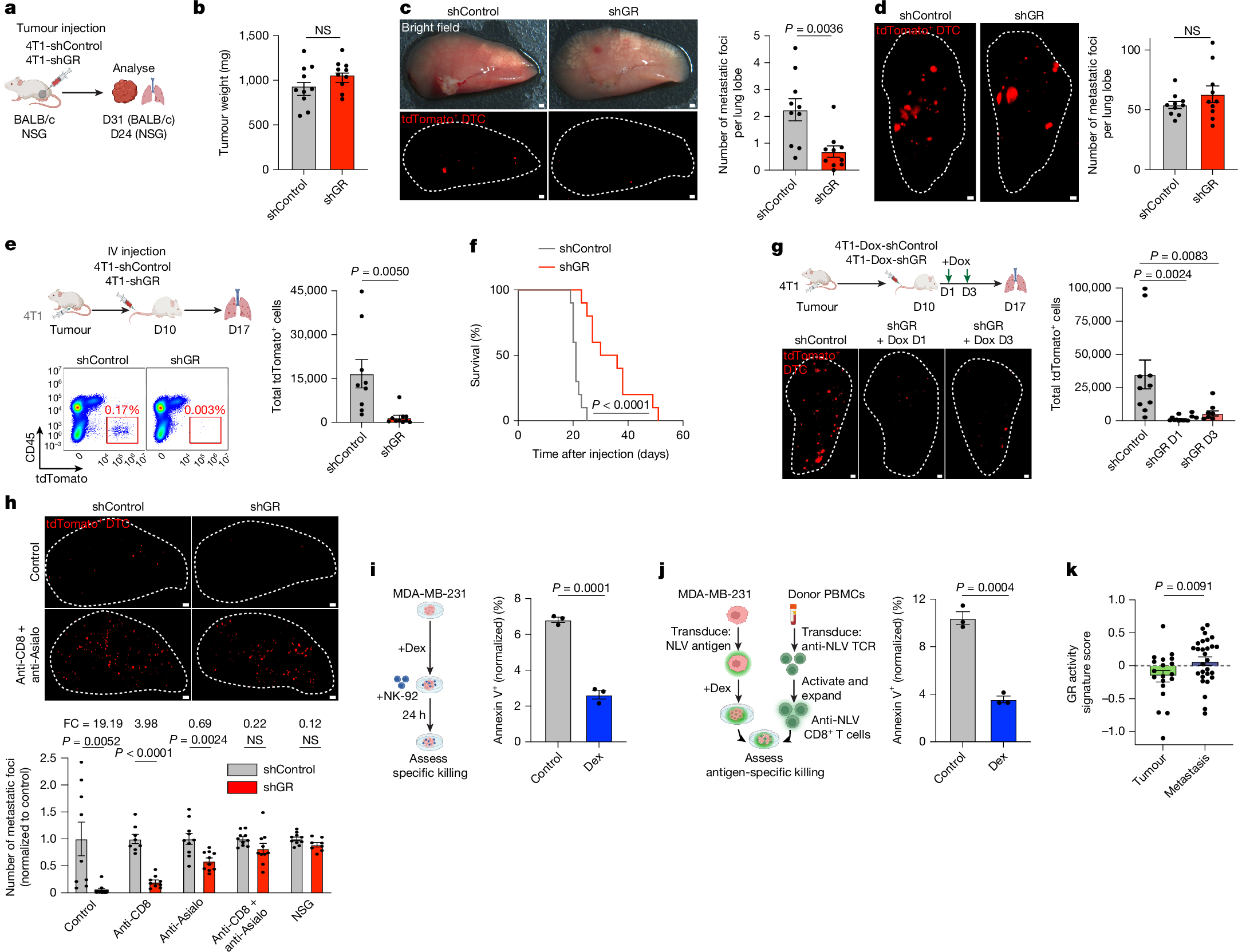
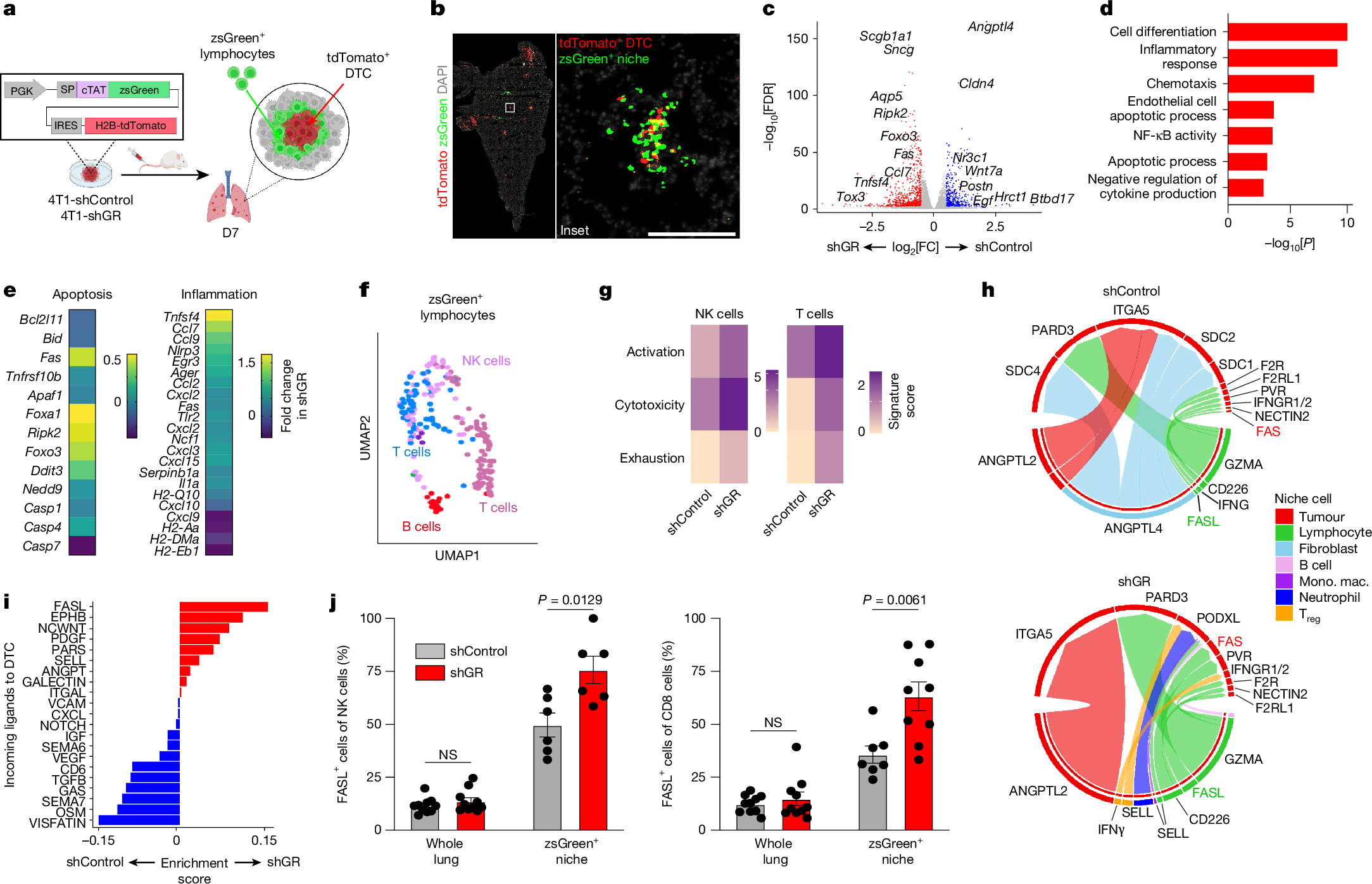
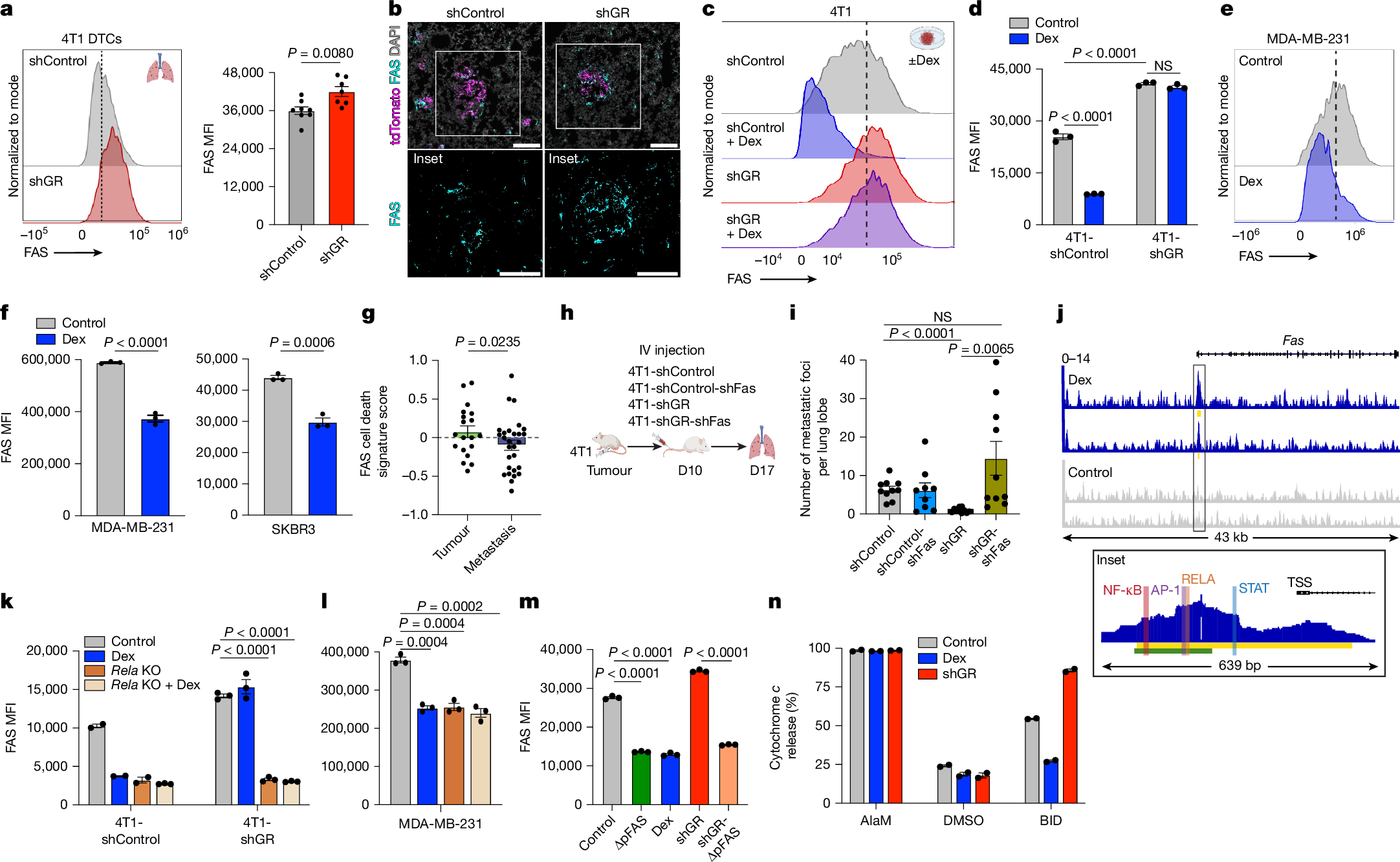
Metastasis is the major cause of death for patients with triple-negative breast cancer and other solid malignancies. Metastases arise from cancer cells that disseminate from the original tumour, survive systemic immune surveillance and colonize new organs1. Little is known about how initial disseminated tumour cells (DTCs) overcome anti-tumour immunity after seeding a new organ. Here we use a visible antigen in a model of triple-negative breast cancer with cognate CD8+ T cells to study the mechanisms of immune evasion in early metastatic seeding. Analysis of surviving DTCs revealed glucocorticoid receptor (GR) activation as a key driver of resistance to both CD8+ T cells and natural killer cells. Niche profiling using an optimized labelling tool identified FAS–FASL as a key pan-cytotoxic pathway against DTCs, which is repressed by GR activation. Pharmacological inhibition of GR in combination with immunotherapy reduced metastatic burden and expanded lifespan in mice. Thus, we identified a mechanism of immune evasion that operates specifically in DTCs, illustrating the unique immune–cancer interactions at this stage in the metastatic cascade. Our findings suggest that there are therapeutic opportunities to eliminate DTCs, separately from treatments aimed at primary tumours, and GR inhibition is one promising target.
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Data availability
The raw data corresponding to bulk RNA-seq datasets generated in this study have been deposited in the Gene Expression Omnibus (GEO) repository under accession numbers GSE313249 (tdTomato+ 4T1 tumour cells from NSG versus BALB/c mice across disease stages) and GSE313241 (GFP+ 4T1 DTCs from Jedi versus NSG mice). scRNA-seq raw data have been deposited in the GEO under accession numbers GSE313245 (GFP+ versus Thy1.1+ 4T1 DTCs after Jedi adoptive transfer) and GSE313243 (tdTomato-cTAT-zsGreen niche labelling of shGR versus shControl 4T1 DTCs). CUT&RUN raw data have been deposited in the GEO under accession number GSE313250. Clinical trial data from TBCRC-030 are available on reasonable request. Any additional data reported in this study are available on request. All other datasets reanalysed in this study are publicly available70 (AURORA US, https://cbioportal.org; TCGA, https://xenabrowser.net/ and http://gepia2.cancer-pku.cn/; meta-analysis of patients with TNBC, https://kmplot.com/analysis; Tabula Muris, https://tabula-muris.sf.czbiohub.org/). Source data are provided with this paper.
Code availability
No custom code was used in the generation of the data in this study. Data analysis using public software packages is described in the Methods.
References
Lambert, A. W., Zhang, Y. & Weinberg, R. A. Cell-intrinsic and microenvironmental determinants of metastatic colonization. Nat. Cell Biol. 26, 687–697 (2024).
Garrido-Castro, A. C., Lin, N. U. & Polyak, K. Insights into molecular classifications of triple-negative breast cancer: improving patient selection for treatment. Cancer Discov. 9, 176–198 (2019).
Pogoda, K., Niwińska, A., Murawska, M. & Pieńkowski, T. Analysis of pattern, time and risk factors influencing recurrence in triple-negative breast cancer patients. Med. Oncol. 30, 388 (2013).
Harper, K. L. et al. Mechanism of early dissemination and metastasis in Her2+ mammary cancer. Nature 540, 588–592 (2016).
Hosseini, H. et al. Early dissemination seeds metastasis in breast cancer. Nature 540, 552–558 (2016).
Hüsemann, Y. et al. Systemic spread is an early step in breast cancer. Cancer Cell 13, 58–68 (2008).