Structural basis of RNA-guided transcription by a dCas12f–σE–RNAP complex
TL;DR
This study reveals a novel RNA-guided transcription mechanism using cryo-EM structures of the dCas12f–σE–RNAP complex. It shows how CRISPR–Cas targeting replaces classical promoter recognition, enabling precise gene activation and expanding programmable transcriptional control.
Key Takeaways
- •The dCas12f–σE system activates gene expression through RNA-guided DNA binding, recruiting RNA polymerase for transcription initiation.
- •CRISPR–Cas targeting largely supplants the classical −35 element recognition in σE-dependent transcription, while the −10 element is stabilized by unusual stacking interactions.
- •This work provides high-resolution structural insights into bacterial gene regulation, opening new avenues for engineered transcriptional control.
Tags
Abstract
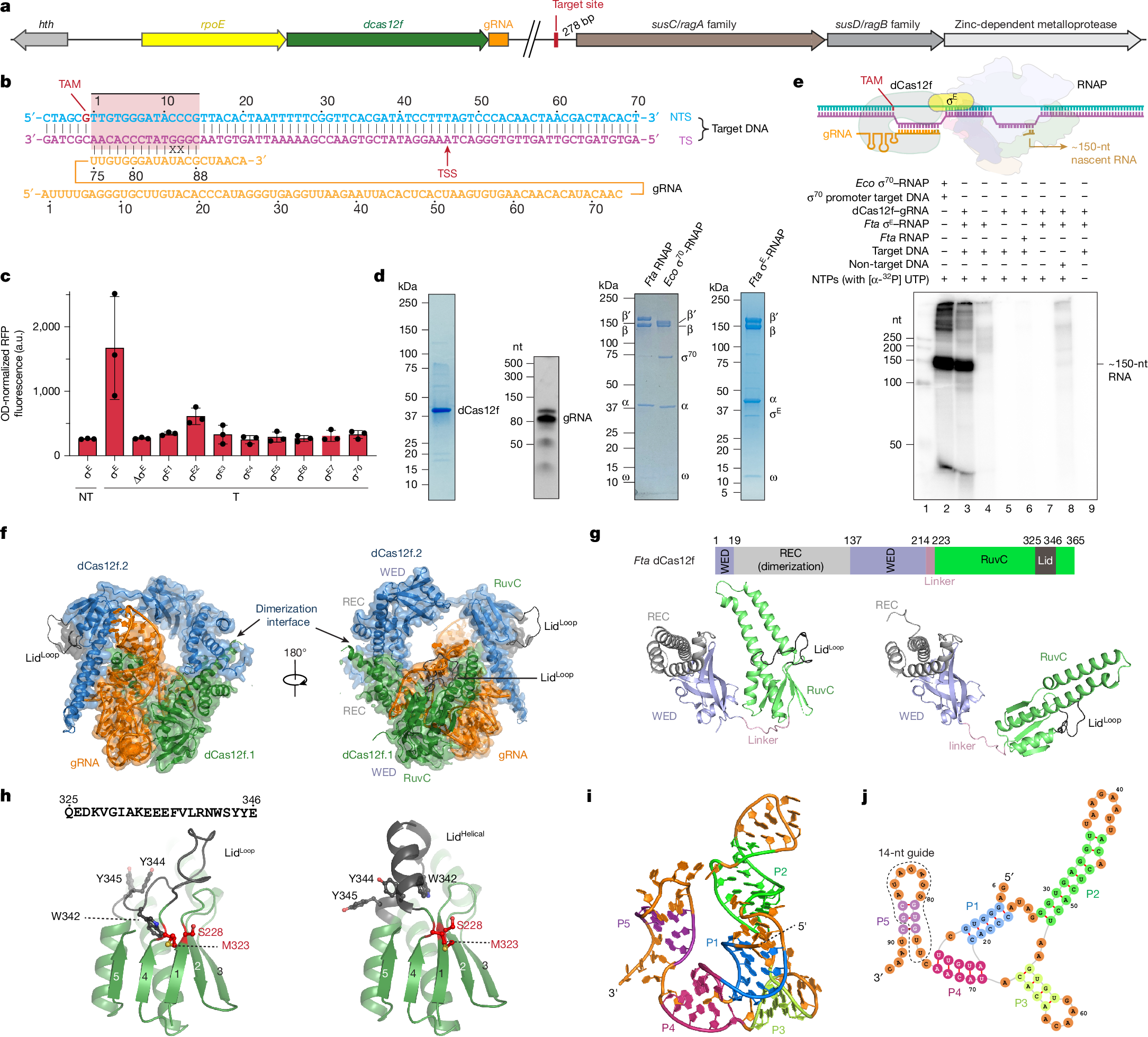
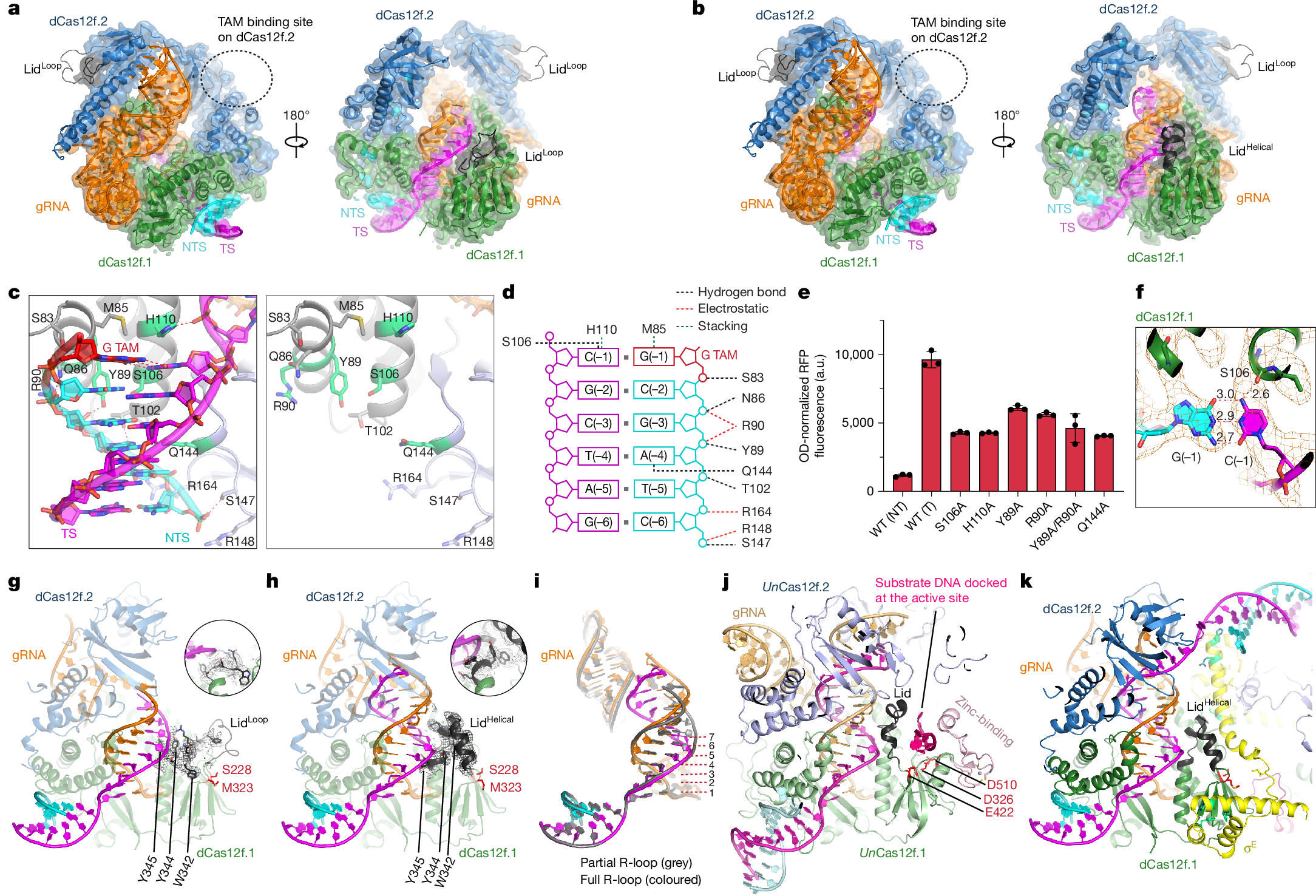
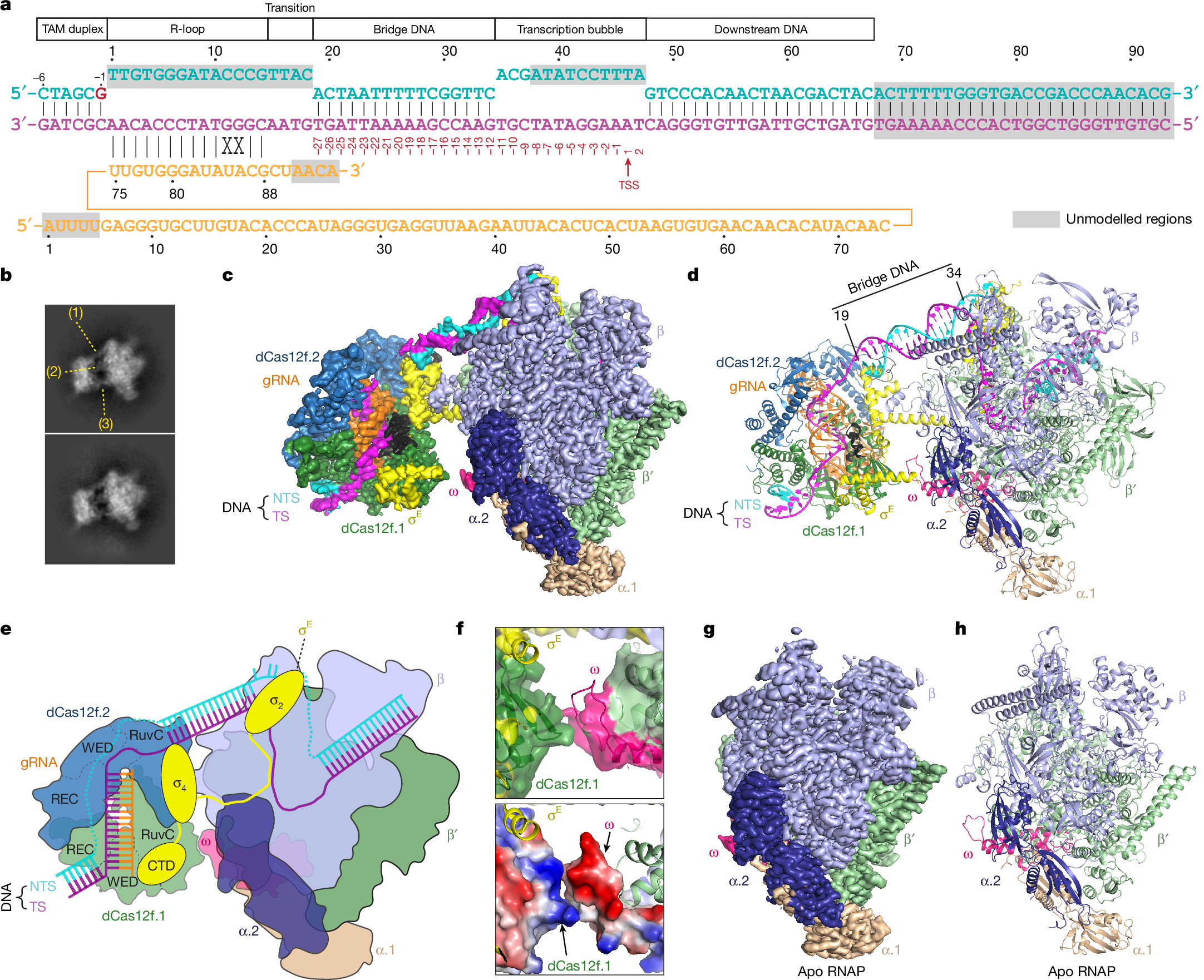
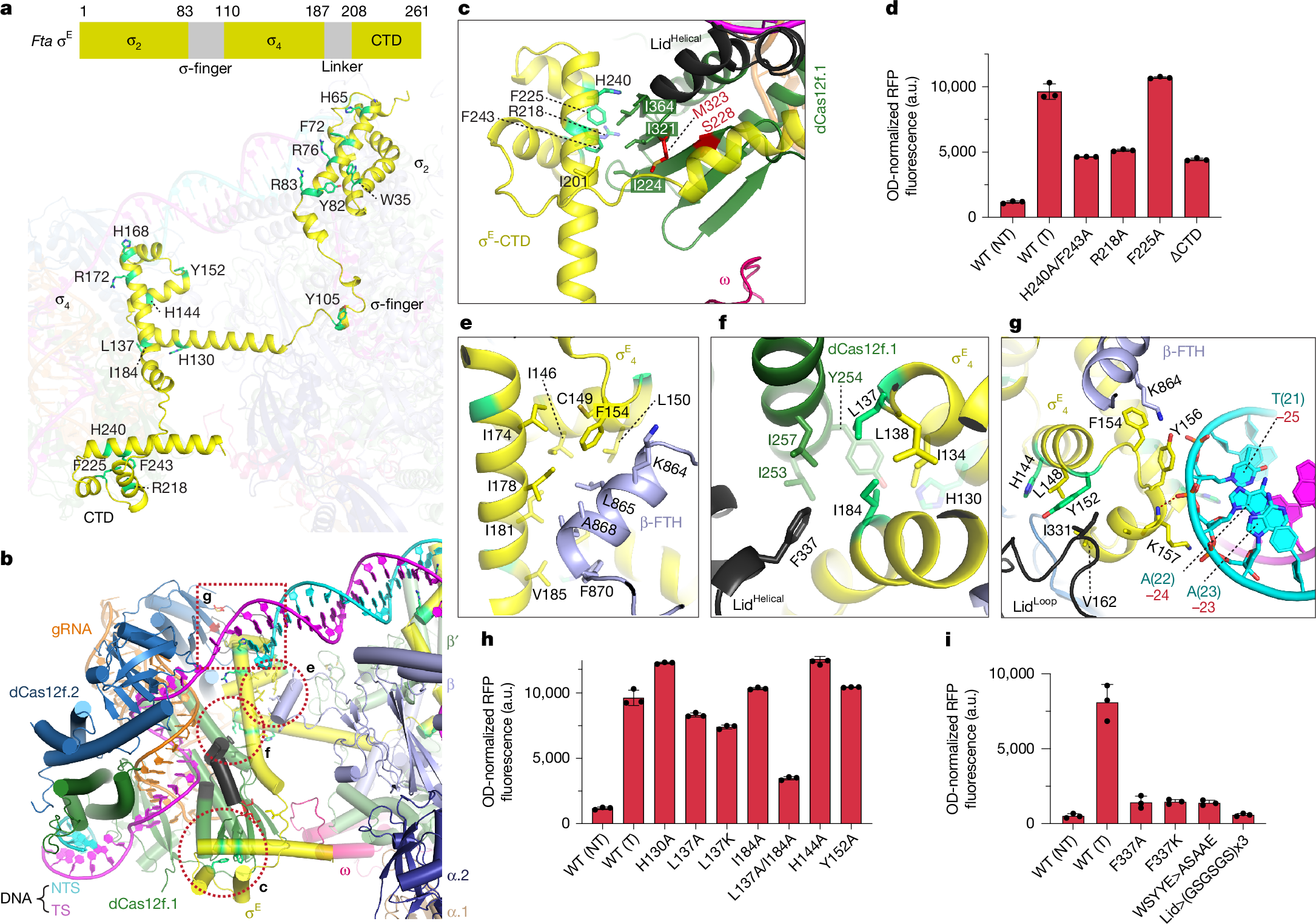
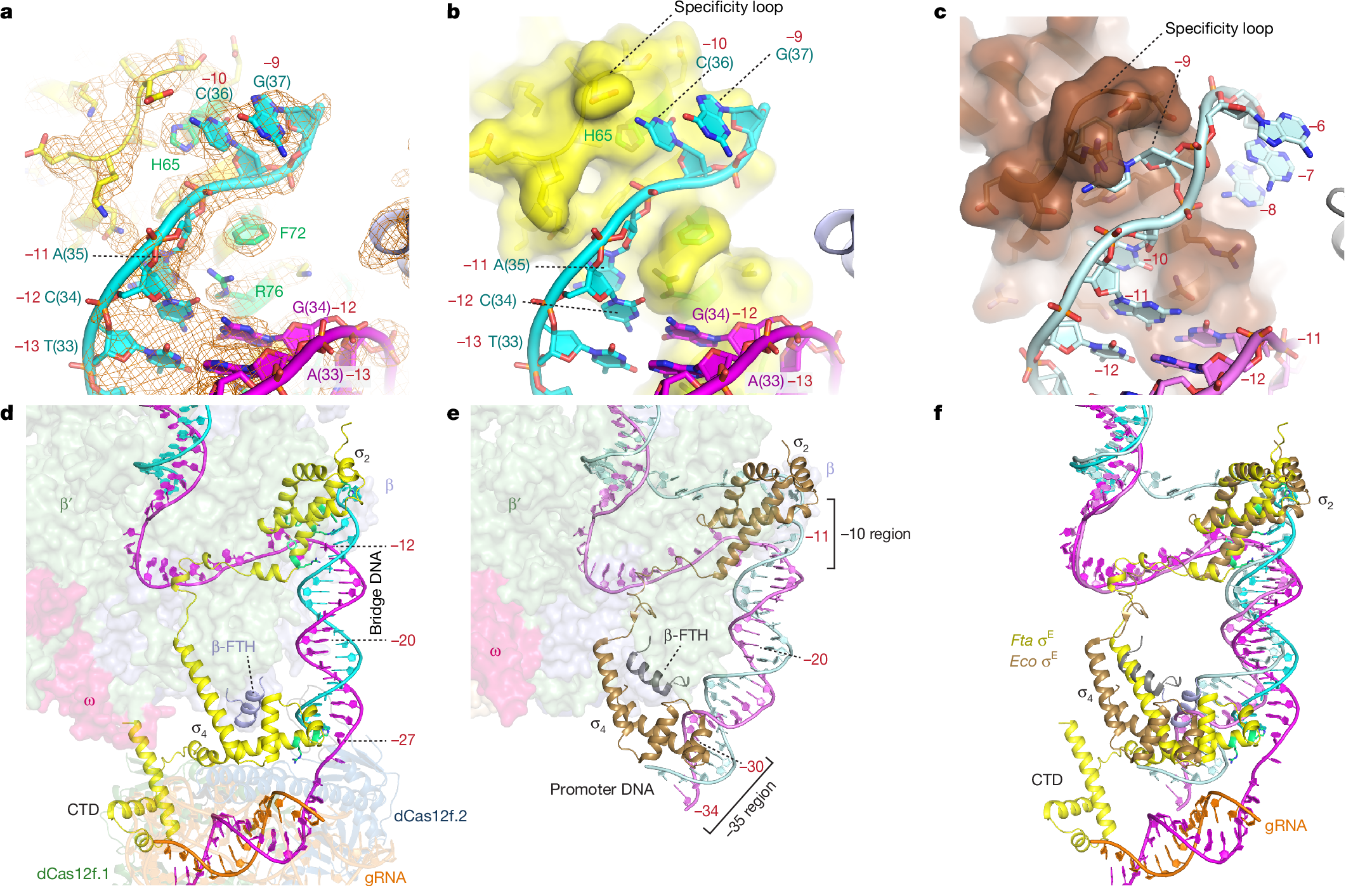
In both natural and engineered biological systems, RNA-guided proteins have emerged as critical transcriptional regulators by modulating RNA polymerase (RNAP) and its associated factors1,2,3. In bacteria, diverse clades of repurposed TnpB and CRISPR-associated proteins repress gene expression by blocking transcription initiation or elongation, enabling non-canonical modes of regulatory control and adaptive immunity1,4,5. A distinct class of nuclease-dead Cas12f homologues (dCas12f) instead activates gene expression through its association with unique extracytoplasmic function sigma factors (σE)6, although the molecular basis has remained elusive. Here we reveal a new mode of RNA-guided transcription initiation by determining the cryo-electron microscopy structures of the dCas12f–σE system from Flagellimonas taeanensis. We captured multiple conformational and compositional states, including the DNA-bound dCas12f–σE–RNAP holoenzyme complex, revealing how RNA-guided DNA binding leads to σE–RNAP recruitment and nascent mRNA synthesis at a precisely defined distance downstream of the R-loop. Rather than following the classical paradigm of σE-dependent promoter recognition, these studies show that recognition of the −35 element is largely supplanted by CRISPR–Cas targeting, whereas the melted −10 element is stabilized through unusual stacking interactions rather than insertion into the typical recognition pocket. Collectively, this work provides high-resolution insights into an unexpected mechanism of RNA-guided transcription, expanding our understanding of bacterial gene regulation and opening new avenues for programmable transcriptional control.
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Data availability
Cryo-EM reconstructions have been deposited in the Electron Microscopy Data Bank under accession nos. EMD-49165, EMD-49173, EMD-49174, EMD-49175 and EMD-49176. Coordinates for atomic models have been deposited in the Protein Data Bank under accession nos. 9N9C, 9N9M, 9N9O, 9N9P and 9N9Q. The raw mass spectrometry data have been submitted to the MassIVE repository under ID MSV000099155. Protein accession numbers for the F. taeanensis σ factor phylogeny are presented in Extended Data Fig. 1. Accession numbers for the RpoZ conservation analysis are listed in Supplementary Data 2. Source data are provided with this paper.
References
Wiegand, T. et al. TnpB homologues exapted from transposons are RNA-guided transcription factors. Nature 631, 439–448 (2024).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Bikard, D. et al. Programmable repression and activation of bacterial gene expression using an engineered CRISPR-Cas system. Nucl. Acids Res. 41, 7429–7437 (2013).
Workman, R. E. et al. A natural single-guide RNA repurposes Cas9 to autoregulate CRISPR-Cas expression. Cell 184, 675–688.e619 (2021).
Wu, W. Y. et al. The miniature CRISPR-Cas12m effector binds DNA to block transcription. Mol. Cell 82, 4487–4502.e4487 (2022).