Mechanism of co-transcriptional cap snatching by influenza polymerase
TL;DR
Cryo-EM structures reveal how influenza polymerase (FluPol) binds to host RNA polymerase II (Pol II) and elongation factor DSIF for cap snatching. FluPol's endonuclease domain positions near Pol II's RNA exit channel, with DSIF stabilizing the interaction, enabling efficient cleavage of capped RNA primers for viral transcription.
Key Takeaways
- •FluPol interacts directly with both Pol II and DSIF, positioning its endonuclease domain near Pol II's RNA exit channel for efficient cap snatching.
- •Phosphorylation of the Pol II CTD is crucial for FluPol recruitment, while DSIF enhances RNA cleavage activity.
- •The cap(1) structure has a minor effect on FluPol binding, but the Pol II–DSIF complex serves as the minimal substrate for cap snatching.
- •Structural insights show that after cleavage, the capped RNA rearranges within FluPol to prime viral transcription initiation.
Tags
Abstract
Influenza virus mRNAs are stable and competent for nuclear export and translation because they receive a 5′ cap(1) structure in a process called cap snatching1. During cap snatching, the viral RNA-dependent RNA polymerase (FluPol) binds to host RNA polymerase II (Pol II) and the emerging transcript2,3. The FluPol endonuclease then cleaves a capped RNA fragment that subsequently acts as a primer for the transcription of viral genes4,5. Here we present the cryogenic electron microscopy structure of FluPol bound to a transcribing Pol II in complex with the elongation factor DSIF in the pre-cleavage state. The structure shows that FluPol directly interacts with both Pol II and DSIF, positioning the FluPol endonuclease domain near the RNA exit channel of Pol II. These interactions are important for the endonuclease activity of FluPol and FluPol activity in cells. A second structure, trapped after cap snatching, shows that the cleaved capped RNA rearranges within FluPol, directing the capped RNA 3′ end toward the FluPol polymerase active site for viral transcription initiation. Together, our results provide the molecular mechanisms of co-transcriptional cap snatching by FluPol.
Main
Influenza is an acute respiratory disease that causes 290,000 to 650,000 human deaths each year6. Influenza is caused by an infection with influenza A or B viruses, which circulate in temperate regions as seasonal influenza6. However, rare zoonotic transmissions can cause pandemic influenza outbreaks with high mortality and economic losses7,8. There is current concern that the unexpected susceptibility of dairy cows to avian H5N1 strains may be a path towards a new pandemic9,10,11. Influenza viruses are segmented negative-sense RNA viruses that infect the respiratory tract epithelial cells in humans8. After infection, the eight viral ribonucleoproteins are released into the cytoplasm and imported into the nucleus, where transcription of viral genes into mRNA and replication of the viral genome occur12,13. Each viral ribonucleoprotein contains a genome segment that is encapsidated by multiple copies of the viral nucleoprotein and one copy of the viral RNA-dependent RNA polymerase (FluPol). FluPol consists of subunits PA, PB1 and PB2 and has been structurally characterized2,14,15.
Viral transcripts must contain a 5′ cap structure and a 3′ poly(A) tail to ensure stability, nuclear export and efficient translation16. However, unlike non-segmented negative-sense RNA viruses, the influenza virus genome does not encode enzymes that synthesize a 5′ cap17. Instead, FluPol utilizes capped RNA primers that are cleaved from nascent host transcripts in a process called cap snatching1,5. The FluPol PB2 cap-binding domain binds a nascent 5′ capped host RNA, and the PA endonuclease domain cleaves off 10–15 nucleotides (nt) from the 5′ end. The 3′-terminal nucleotides of this RNA primer then anneal to the 3′ end of the viral genome segment and prime transcription of the viral mRNA14,18,19.
Capped host transcripts are synthesized by cellular Pol II. Pol II transcription starts with assembling a pre-initiation complex consisting of Pol II and the general transcription factors at gene promoters20. To escape from the gene promoter, the largest Pol II subunit RPB1 C-terminal domain (CTD) heptad repeats are phosphorylated at serine 5 and serine 7 by the TFIIH CDK-activating kinase (CAK)21,22. CTD phosphorylation and the growing nascent RNA transcript cause the initiation factors to dissociate from Pol II22,23. Recruitment of the elongation factor DSIF after synthesis of around 20 nt of RNA establishes the early Pol II elongation complex (Pol II–DSIF). This complex is then converted to a paused elongation complex (PEC) containing the negative elongation factor NELF at a transcript length of 25–50 nt (refs. 23,24,25). Synthesis of the 5′ cap occurs co-transcriptionally by the capping enzymes RNGTT, RNMT and CMTR1 (ref. 25) in the context of the Pol II–DSIF elongation complex or the PEC. RNGTT is a bifunctional enzyme that acts as a triphosphatase and guanylyltransferase, creating a GpppN structure at the 5′ end of the Pol II transcript. RNMT and CMTR1 are methyltransferases that add a methyl group to N7 of the cap guanosine and the 2′-OH of the first regular nucleotide, respectively, producing the m7GpppmN cap(1) structure25, which the cap-binding domain of FluPol subunit PB2 tightly binds during cap snatching26,27.
Cap snatching depends on host transcription, as it has been shown that inhibition of Pol II using α-amanitin impairs viral replication3. FluPol localizes primarily at the 5′ end of host genes and associates with the Pol II CTD that is phosphorylated at serine 5 residues, indicating that cap snatching occurs during early phases of Pol II transcription2,28,29,30. Cell-based protein–protein interaction assays indicate that FluPol binds not only to the CTD but also to the Pol II body31. Co-immunoprecipitation–mass spectrometry experiments have shown that the elongation factor DSIF co-purifies with FluPol5,32, and other studies suggest that FluPol depends on the cap(1) structure for cap snatching26. However, how FluPol interacts with the host transcription machinery for cap snatching at the molecular level is unknown.
Here we show that FluPol binds to the transcribing Pol II–DSIF complex for efficient cap snatching. Furthermore, we report two cryogenic electron microscopy (cryo-EM) structures of FluPol bound to a Pol II–DSIF elongation complex before and after endonucleolytic RNA cleavage by FluPol. The structures show that during cap snatching, the PA endonuclease domain of FluPol binds near the RNA exit channel of Pol II and that this interaction is stabilized by DSIF. Furthermore, using cell-based minigenome assays, we confirm that mutation of residues forming the interface between FluPol and the Pol II–DSIF elongation complex reduces FluPol activity in cells. In summary, we present the molecular mechanism of cap snatching by FluPol.
FluPol snatches cap from Pol II elongation complex
To study the molecular basis of cap snatching, we first investigated how the formation of a complex between FluPol and transcribing Pol II (Pol II elongation complex) depends on the cap(1) structure and CTD phosphorylation. We purified Sus scrofa Pol II (99.9% sequence identity to human Pol II, with 4 amino acid differences) from the endogenous source33. Whereas in preliminary studies reconstituting the cap-snatching complex30, we used bat FluPol(H17N10), here we used recombinant, viral promoter-bound FluPol from the influenza strain A/Zhejiang/DTID-ZJU01/2013(H7N9)34,35 (Extended Data Fig. 1a). To reduce RNA cleavage and enhance complex stability, we used the PA(E119D) mutant of FluPol (FluPolE119D; Extended Data Fig. 3j), which has impaired endonuclease activity36,37. A Pol II elongation complex containing a 35-nt cap(1)-RNA, 45-nt template, and non-template DNA was assembled as established previously38. The 35-nt RNA length was chosen considering a 12-nt RNA primer produced by cap snatching19,39, an additional 3 nt bound by the PA endonuclease37, and 20 nt RNA bound within the Pol II elongation complex33.
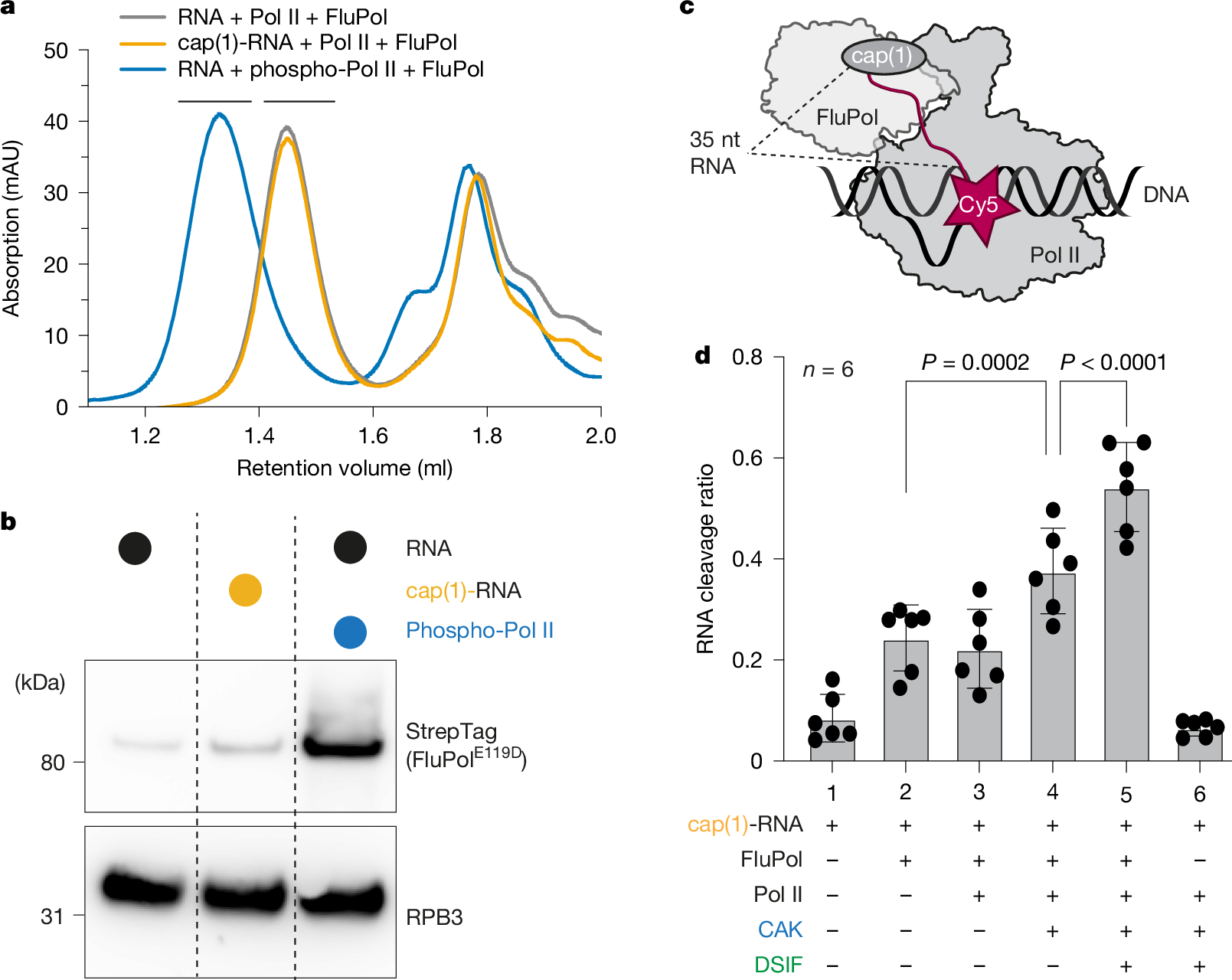
We next monitored binding of FluPolE119D to the Pol II elongation complex by size-exclusion chromatography (SEC) using unmodified RNA and Pol II, cap(1)-RNA, or Pol II that was phosphorylated with CAK. Without CTD phosphorylation and a cap(1) structure, co-elution of FluPol with Pol II could barely be detected (Fig. 1a,b). When a cap(1)-modified RNA was used, the signal for FluPol in the Pol II containing peak slightly increased (Fig. 1b). However, when the Pol II CTD was phosphorylated by CAK, the amount of FluPol associated with Pol II in the peak fractions strongly increased (Fig. 1b). Additionally, the elution volume of the complex peak shifted towards higher molecular weight, indicating the formation of a stable complex (Fig. 1a). Thus, the addition of a cap(1) structure to the RNA has a negligible effect on the interaction between FluPol and the Pol II elongation complex. By contrast, phosphorylation of the Pol II CTD is the main determinant for the recruitment of FluPol to a Pol II elongation complex, consistent with in vivo data demonstrating the importance of the Pol II CTD for viral transcription2,29.
a, Absorbance at 280 nm of analytical SEC runs of Pol II elongation complex containing a 35-nt RNA with or without cap(1) and with or without CAK phosphorylation, and with FluPolE119D. Different colours represent different chromatography runs. Black bars above the chromatogram depict Pol II complex fractions that were analysed by Western blot in b. b, Western blot of Pol II containing peak fractions stained against RPB3 (Pol II) and Twin-StrepTag (FluPol subunit PB2). Individual lanes represent different SEC runs. c, Schematic of the endonuclease cleavage assay. The cap(1)-RNA is Cy5-labelled on the 3′ end. d, The fraction of cleaved RNA (intensity of cleaved product divided by intensity of all bands, see Extended Data Fig. 1c) depends on the factors added. Each point reflects one experimental replicate (n = 6); mean ± s.d. P values were calculated using a linear mixed-effects model (substrate as a fixed effect, experimental replicate as a random effect, two-sided, no multiple testing correction).
We next tested whether the increased affinity of FluPolE119D to Pol II by CTD phosphorylation also results in enhanced endonuclease activity by wild-type FluPol. To monitor RNA cleavage, we developed a fluorescence-based assay using in vitro-capped RNA labelled with Cy5 at the 3′ end (Fig. 1c). We assembled Pol II elongation complexes in vitro essentially as described above, added wild-type FluPol and then visualized the cleaved 3′ end of the RNA that remains attached to Pol II by denaturing PAGE. The primary cleavage product detected was 20–25 nt long, corresponding to the expected 10- to 15-nt primer generated by FluPol (Extended Data Fig. 1b). Additionally, small amounts of an additional cleavage product of around 30 nt were produced. Comparing the different RNA substrates, we did not observe an increase in RNA cleavage by FluPol in the context of a Pol II elongation complex compared to free RNA (Fig. 1d, Extended Data Fig. 1b,c and Supplementary Table 1). However, RNA cleavage increased when we phosphorylated the CTD of Pol II by adding CAK (Fig. 1d and Extended Data Fig. 1b), in line with previous reports40. This suggests that CTD phosphorylation enhances recruitment of FluPol to Pol II and stimulates cleavage of RNA that is bound to Pol II.
Next, we tested whether the presence of the elongation factor DSIF, which binds Pol II during early elongation, stimulates the cleavage of Pol II-bound RNA. Indeed, cleavage of Pol II-bound RNA was stimulated ~1.5-fold when DSIF was added in excess to the Pol II (Fig. 1d and Extended Data Fig. 1b, c). Finally, we tested whether wild-type FluPol can extend the snatched RNA primer using a radioactive FluPol RNA extension assay. We found that FluPol alone can extend the cleaved RNA fragments to some degree. Furthermore, FluPol-dependent RNA extension increases in the presence of a Pol II–DSIF elongation complex, which indicates that the more efficient endonuclease reaction provides more usable RNA primers for FluPol transcription (Extended Data Fig. 1d).
In summary, the cap(1) structure only has a minor effect on FluPol binding to Pol II, whereas CTD phosphorylation by CAK strongly enhances FluPol recruitment and stimulates cleavage of Pol II-bound RNA by FluPol to some extent. Additionally, DSIF, when added to the Pol II elongation complex, significantly enhances RNA cleavage further, suggesting that DSIF is part of the Pol II complex that is recognized by FluPol. Moreover, we have demonstrated that the RNA emerging from the Pol II–DSIF elongation complex can be used to prime RNA synthesis by FluPol. Thus, we conclude that FluPol recognizes the phosphorylated Pol II–DSIF elongation complex as a minimal substrate for efficient cap snatching.
FluPol–Pol II–DSIF complex structure
After determining the components required for efficient cap snatching by FluPol in vitro, we next sought to structurally characterize a cap-snatching complex comprising FluPol, Pol II, DSIF and capped RNA by cryo-EM. To that end, we first assembled a Pol II–DSIF elongation complex containing a 35-nt cap(1)-RNA in the presence of the CAK and ATP to phosphorylate the Pol II CTD. To capture the normally transient cap-snatching complex prior to RNA cleavage, we added FluPolE119D at low Mg2+ concentration, a condition in which cleavage is minimal (Extended Data Fig. 1e). The complex was purified and stabilized using GraFix41 prior to cryo-EM sample preparation (Extended Data Fig. 2a). Cryo-EM data acquisition yielded 6,423,874 particles that were further sorted by 3D classification, which yielded a subset of 369,858 particles that show good density for the Pol II–DSIF elongation complex, as well as FluPol resolved at 3.3 Å overall resolution (Extended Data Fig. 2b–h and Extended Data Table 1). From this consensus refinement, we performed focused refinements of FluPol and the Pol II–DSIF elongation complex (with resolutions of 2.90 Å and 2.94 Å, respectively), which enabled us to build and refine an atomic model for the complete cap-snatching complex (Fig. 2a).
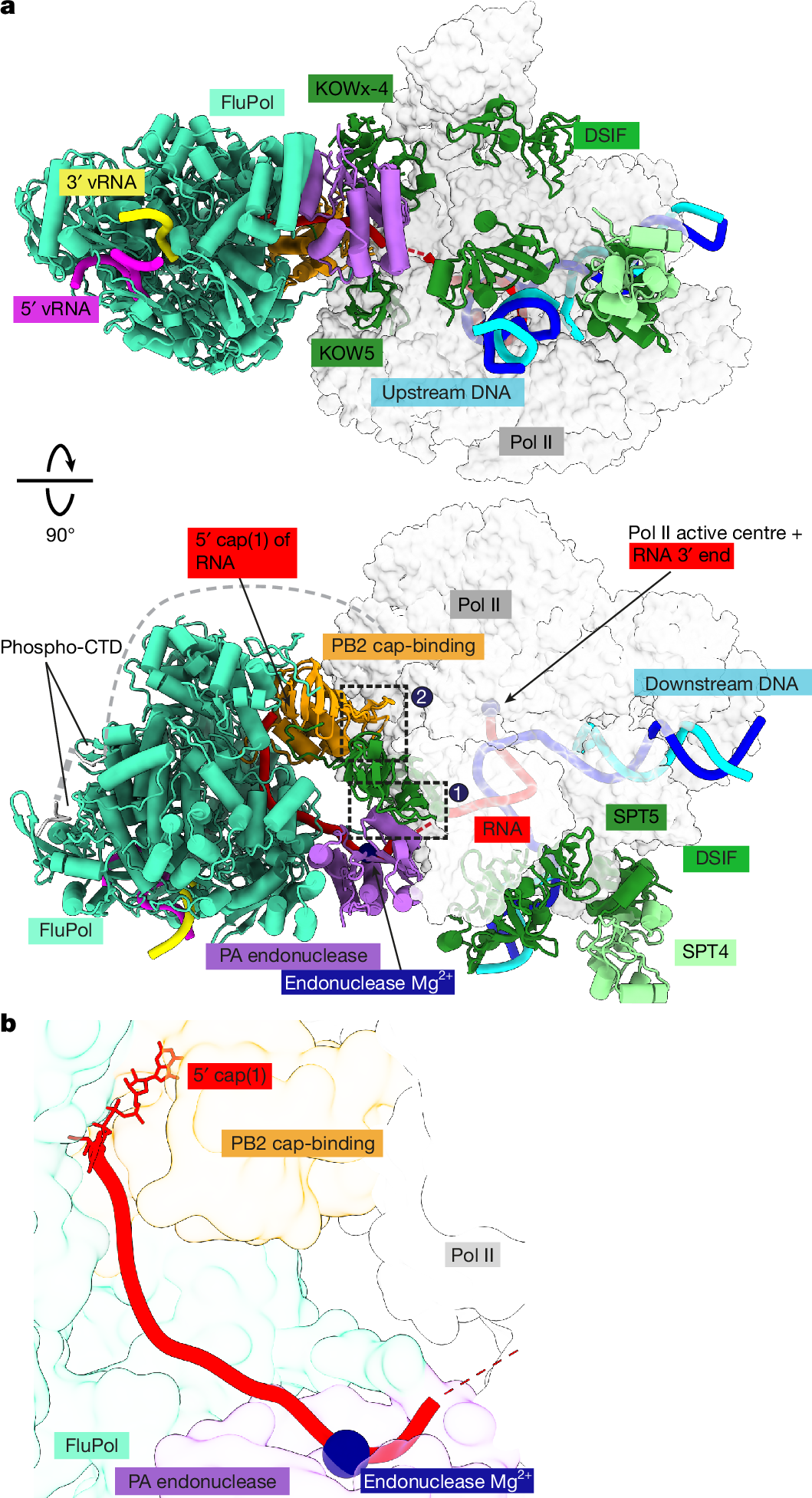
a, Two views of the overall structure of the pre-cleavage FluPol–Pol II–DSIF elongation complex complex in cartoon representation except Pol II, which is shown as surface. Dashed black boxes represent the locations of the two interfaces shown in Fig. 3a–c. The structure is shown in a FluPol side view and Pol II top view (top) as well as front view of FluPol and side view of Pol II (bottom). b, The RNA path within FluPol. Proteins are shown as transparent surfaces. The RNA is shown as ribbon tracing of the backbone. Parts of the FluPol model were removed for clarity.
The structure shows that FluPol binds to the Pol II–DSIF elongation complex near the RNA exit channel of Pol II (Fig. 2a). The PA endonuclease of FluPol interacts with the KOWx-4 domain of DSIF that forms a clamp around the exiting RNA in the absence of FluPol33 (Fig. 2a, interface 1). In the complex, KOWx-4 is rotated approximately 180° around its longitudinal axis and shifted by around 22 Å compared with the Pol II–DSIF elongation complex structure33, and the Pol II stalk containing subunits RPB4 and RPB7 is also repositioned (Extended Data Fig. 3a,b). The PB2 cap-binding domain of FluPol inserts between the Pol II subunits RPB1, RPB3 and RPB11 to bind the Pol II dock domain, which is located below the RNA exit channel of Pol II (Fig. 2a, interface 2). In line with our observation that FluPol recruitment to Pol II strongly depends on CTD phosphorylation, we observe density for serine 5 phosphorylated CTD residues in the two previously reported CTD binding sites of FluPol2,