DICER cleavage fidelity is governed by 5′-end binding pockets
TL;DR
DICER's cleavage fidelity is regulated by dual 5′-end binding pockets: a G-favored pocket promoting precise cleavage at DC21 and a U-favored pocket favoring DC22. This mechanism integrates RNA motifs and domain motions for accurate small-RNA biogenesis.
Key Takeaways
- •DICER uses two distinct 5′-end binding pockets: a G-favored pocket that enhances cleavage accuracy at DC21 and a U-favored pocket that promotes cleavage at DC22.
- •5′-G nucleotides, contrary to previous beliefs, improve cleavage precision for many substrates, as shown through massively parallel dicing assays and cryo-EM structures.
- •RNA motifs like mWCU and YCR cooperate with 5′-end binding rules to refine cleavage specificity, with conflicts triggering conformational adjustments in RNA and DICER domains.
- •Dynamic rearrangements of the dsRBD and PAZ domains help align RNA substrates with catalytic centers, ensuring precise double-strand cleavage in the dicing state.
- •This dual-pocket mechanism advances understanding of microRNA biogenesis by integrating end-binding preferences, RNA features, and domain motions for accurate RNA processing.
Tags
Abstract
RNA interference (RNAi) depends on DICER, an essential enzyme that processes RNA precursors into small regulatory RNAs. DICER cleaves RNA precursors according to the 5′-end counting rule, in which RNA length is measured from the 5′-end1,2,3. Previous work proposed a single 5′-end binding pocket that disfavours guanosine (5′-G), leading to cleavage inaccuracies4. Here we show that 5′-G promotes precise cleavage for many substrates. Using massively parallel dicing assays and cryo-electron microscopy, we identify a conserved guanosine-favoured (G-favoured) binding pocket in DICER, distinct from the previously described uridine-favoured (U-favoured) pocket. Together, these pockets influence the alignment between 21-nucleotide and 22-nucleotide cleavage registers, expanding the mechanism of small-RNA biogenesis in metazoan DICERs. We also find that conflicts between 5′-end binding and RNA-motif recognition can trigger RNA conformational adjustments that preserve accurate cleavage-site selection. In addition, conformational adjustments of the double-stranded RNA-binding domain (dsRBD) and PAZ domain help to align substrates with the catalytic centres for precise double-strand cleavage. These results show that the DICER cleavage mechanism integrates dual 5′-end binding pockets, RNA-motif influence and domain motions, advancing our understanding of microRNA biogenesis.
Main
DICER is an evolutionarily conserved RNase III enzyme that has a central role in RNA silencing, a crucial gene regulatory mechanism that influences diverse biological processes. DICER processes precursor microRNAs (pre-miRNAs) and long double-stranded RNAs (dsRNAs) into small regulatory RNAs, microRNAs (miRNAs) and small interfering RNAs (siRNAs), which target mRNAs for degradation or translational repression through the RNA-induced silencing complex (RISC)1,2,3. This cleavage activity also drives short-hairpin RNA (shRNA) technology, in which synthetic shRNAs are processed by DICER into siRNA duplexes that mimic endogenous RNAi pathways5,6,7,8,9,10.
Over the past two decades, studies have elucidated key aspects of DICER’s cleavage mechanism, including 5′- and 3′-end counting rules that determine cleavage sites around 21–22 nucleotides (nt) from the RNA ends4,11,12,13,14,15,16,17. Structural studies have shown that the 3′-end binds within a conserved pocket in DICER’s PAZ domain, whereas the 5′-end interacts with a pocket in DICER’s platform domain4,14,17,18,19,20,21,22,23,24,25. In human, mouse and fly DICER enzymes, the 3′-end terminal phosphate interacts with a cluster of conserved tyrosines and basic residues4,17,18,19,21,22. Similarly, plant DICER-like enzymes (for example, Arabidopsis thaliana DCL1 and DCL3) have a conserved 3′-end docking pocket, highlighted by aligned conserved tyrosines, basic residues, and phenylalanine24,25. These findings suggest that the 3′-end binding pocket is conserved across DICER and DICER-like enzymes.
The previous effort to identify the 5′-end binding pocket in DICER used crystal structures of its PAZ–platform–connector helix cassette bound to siRNAs, revealing a phosphate-binding pocket17. However, these structures might not represent the dicing state. A cryo-electron microscopy (cryo-EM) study resolved DICER in the dicing state, detailing a 5′-end binding pocket for 5′-U, involving R790, R821 and R10034. This pocket disfavours guanosine (5′-G), owing to steric clashes with the arginine residue (R821), suggesting that 5′-G reduces cleavage accuracy4. Similarly, a dicing-state structure of fly Dcr-1 identified a 5′-end binding pocket for 5′-U, with R1027 (conserved with R821 in human DICER) located in this pocket21. In A. thaliana DCL3, the 5′-end binding pocket consists of histidine and arginine residues, which are not conserved with human DICER25. By contrast, the 5′-end binding pockets of mouse DICER, fly Dcr-2 and Arabidopsis thaliana DCL1 remain undefined19,22,23,24. These findings suggest that, unlike the conserved 3′-end binding pocket, the 5′-end binding pocket varies among enzymes and is challenging to determine. Moreover, structural studies of human DICER and fly Dcr-1 have relied on RNA substrates with fixed 5′-U, and the effects of diverse RNA ends have not been examined.
DICER uses fine-tuning mechanisms, such as sequence motifs (for example, mWCU, YCR and GYM), bulges, loops and mismatches, to achieve precise cleavage at either position 21 (DC21) or position 22 (DC22) from the 5′-end6,9,26,27,28,29,30,31,32,33,34,35,36,37. However, the interaction between these features and end-binding rules, as well as how DICER coordinates RNA motifs with end-binding preferences for precise cleavage, remain unclear.
In this study, using massively parallel dicing assays and single-particle cryo-EM, we show that 5′-G, contrary to previous reports, enhances cleavage precision at DC21 for many substrates. Cryo-EM reconstructions uncover a previously unrecognized G-favoured binding pocket, distinct from the U-favoured binding pocket, directing cleavage to DC22. This dual-pocket mechanism reshapes our understanding of small-RNA biogenesis, showing how DICER integrates end-binding rules with RNA features for precise cleavage. RNA motifs such as mWCU and YCR cooperate with the 5′-end binding rule to refine specificity, and conflicts induce RNA conformational changes that override end-binding preferences. Cryo-EM further reveals dynamic rearrangements of the dsRBD and PAZ domains, repositioning RNA for precise catalysis and providing a framework for RNA processing in silencing pathways.
DICER cleavage accuracy is enhanced by 5′-G
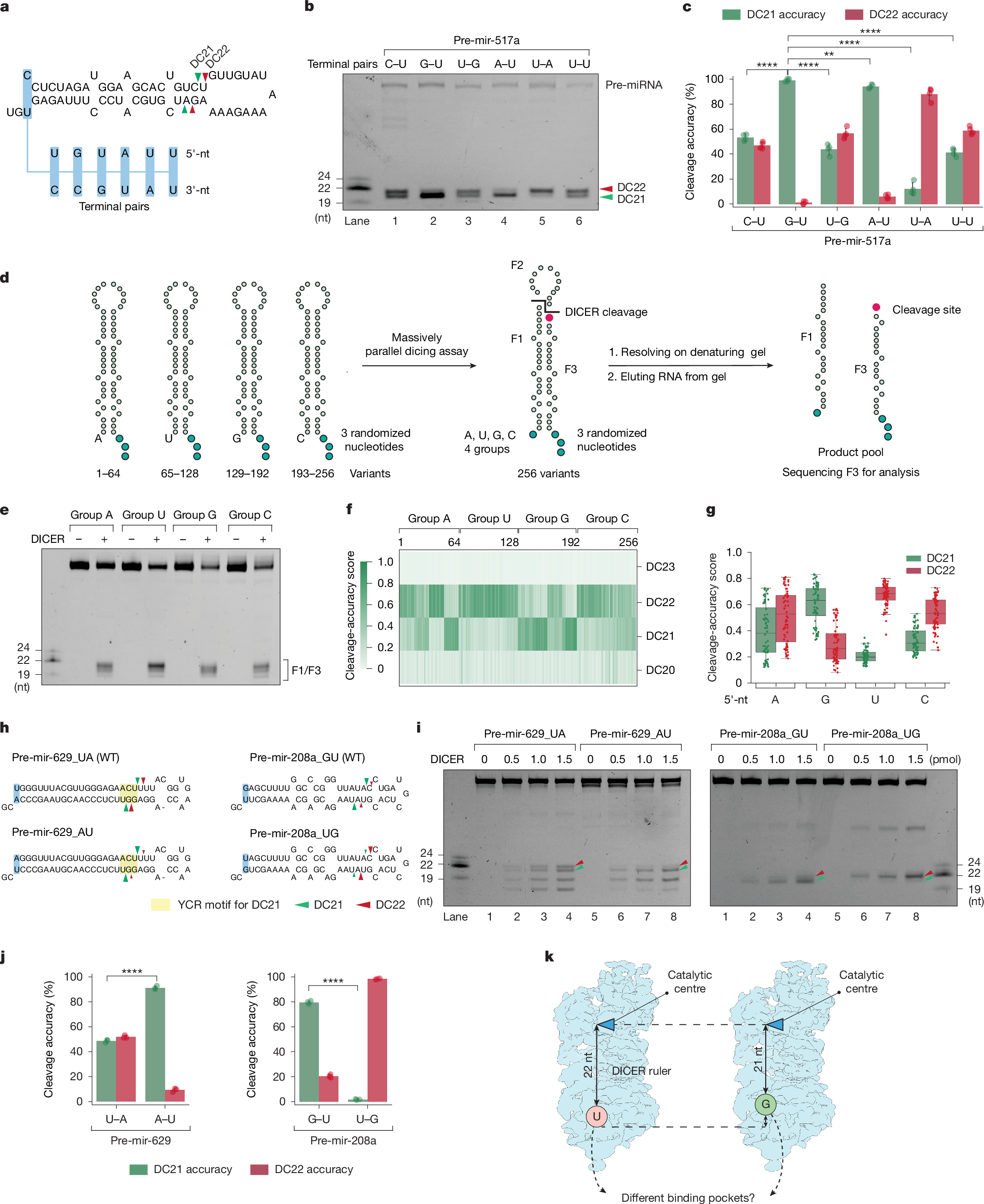
When testing pre-mir-517a, a pre-miRNA with a 5′-C (pre-mir-517a_CU), we observed cleavage at both the DC21 and the DC22 site (Fig. 1a–c). To assess the contribution of the 5′-nucleotide (5′-nt), we replaced the 5′-C with other nucleotides and examined dicing outcomes. Substituting 5′-C with 5′-A (pre-mir-517a_AU) or 5′-G (pre-mir-517a_GU) shifted cleavage predominantly to DC21, with 5′-G showing the highest accuracy, whereas 5′-U (pre-mir-517a_UU), like 5′-C, supported cleavage at both sites (Fig. 1b,c and Extended Data Fig. 1a). Because 5′-G and 5′-A pair with U on the 3′-strand, whereas 5′-C and 5′-U do not, these substitutions altered the overhang geometry. To isolate the effect of the 5′-nt from the overhang context, we tested pre-mir-517a variants with comparable overhangs. Notably, G–U (pre-mir-517a_GU) showed higher DC21 accuracy than U–G, whereas A–U favoured DC21 and U–A favoured DC22 (Fig. 1b,c). These findings refute the hypothesis that 5′-G reduces cleavage accuracy4, and reveal that the 5′-nt strongly influences DICER’s cleavage-site preference: 5′-G and 5′-A promote DC21, whereas 5′-C and 5′-U favour DC22.
a,Schematic of pre-mir-517a substrates, illustrating terminal pairs (for example, G–U and C–U) at the 5′- and 3′-ends. Green and red arrowheads indicate DC21 and DC22 cleavage sites, located 21 and 22 nt from the 5′-end, producing fragments F1, F2 and F3. b, In vitro dicing assays for pre-mir-517a and variants using 2 pmol RNA and 1 pmol DICER. F1 and F3 fragments are 22 nt (DC22) or 21 nt (DC21), whereas F2 is undetectable on gels. c, Cleavage accuracy of DICER at DC21 and DC22 for pre-mir-517a variants, based on four independent experiments. Statistical significance was determined using a two-tailed, two-sample t-test (**P < 0.01, ****P < 0.0001). d, Schematic of massively parallel dicing assays using pre-mir-324 groups with fixed 5′-nt (A, G, U or C) and randomized 3′-overhang. Cleaved products (F3) were cloned, sequenced and analysed. e, Cleavage patterns from pre-mir-324 groups with distinct 5′-end nucleotides. f, Cleavage sites (DC20–DC23) identified from sequencing results. n = 256 variants. g, Cleavage accuracy at DC21 and DC22 for pre-mir-324 groups, calculated from sequencing data. n = 64 for each comparing group. h, Diagrams of pre-mir-629, pre-mir-208a and variants, showing terminal pairs. Green and red arrowheads indicate DC21 and DC22 cleavage sites. i, In vitro dicing assays for pre-mir-629, pre-mir-208a and variants using 2 pmol RNA and 1 pmol DICER. F1 and F3 fragments are 22 nt (DC22) or 21 nt (DC21); F2 is undetectable. Green arrowheads, DC21; red arrowheads, DC22. j, Cleavage accuracy of DICER at DC21 and DC22 for pre-mir-629 and pre-mir-208a variants, based on three independent experiments. Statistics as in c. k, Mechanistic model illustrating how 5′-nt affects DICER cleavage. Guanine (G) and adenine (A) favour DC21 cleavage, whereas uracil (U) and cytosine (C)—especially U—favour DC22 cleavage.
Parallel dicing assays confirm the effects of the 5′-nt
To generalize these findings, we performed massively parallel dicing assays using pre-mir-324, cleaved by DICER at DC20, DC21 and DC2238,39,40. We synthesized four groups of pre-mir-324 variants, each with a specific 5′-nt and randomized 3′-ends, producing 64 unique sequences per group (Fig. 1d). After DICER cleavage, F3 fragments were sequenced to identify cleavage sites (Fig. 1e and Extended Data Fig. 1b). The assays were highly reproducible, with strong correlation between replicates (Extended Data Fig. 1c,d). Sequencing confirmed that DICER cleaves pre-mir-324 mainly at DC21 and DC22 (Fig. 1f). Whereas the 3′-nt had no detectable effect in our randomized pre-mir-324 context (Extended Data Fig. 1e), the 5′-nt strongly influenced site preference: 5′-G yielded the highest DC21 accuracy, followed by 5′-A, whereas 5′-C and 5′-U favoured DC22 (Fig. 1g). These results confirm that 5′-G enhances, rather than impairs, cleavage accuracy by promoting DC21 cleavage, underscoring the prominent role of the 5′-nt in determining DICER’s cleavage-site preference.
Effects of the 5′-nt validated across pre-miRNAs
To validate the role of the 5′-nt in cleavage specificity, we synthesized pre-mir-324 variants with a fixed 3′-end and varying 5′-nts (Extended Data Fig. 1f). Consistent with the massively parallel assays, 5′-G and 5′-U promote cleavage at DC21 and DC22, respectively, whereas 5′-A and 5′-C support cleavage at both sites, with slight preferences for DC21 and DC22, respectively (Extended Data Fig. 1g,h). To isolate the influence of the 5′-nt, we generated variants with similar 2-nt overhang structures but differing base pairs (A–U versus U–A, or G–U versus U–G), confirming that 5′-G strongly favours DC21, 5′-U favours DC22 and 5′-A promotes DC21 more than 5′-U but less than 5′-G (Extended Data Fig. 1i–k). Finally, variants with similar 3′-nt overhangs but differing 5′-nts showed that 5′-A favoured DC21, whereas 5′-U and 5′-C supported DC22, with 5′-U showing the strongest bias (Extended Data Fig. 1l–n).
To test whether the effects of the 5′-nt apply to other pre-miRNAs, we examined pre-mir-629 and pre-mir-208a, which naturally have 5′-U and 5′-G, respectively (Fig. 1h). In pre-mir-629, DICER cleaves at both DC21 and DC22, with DC21 cleavage driven by a YCR motif. Changing 5′-U to 5′-A reduces DC22 cleavage, confirming that 5′-U enhances DC22 cleavage (Fig. 1i,j). Similarly, pre-mir-208a shows DC21 cleavage with 5′-G and DC22 cleavage with 5′-U (Fig. 1i,j). These findings show that 5′-G promotes DC21, whereas 5′-U promotes DC22 (Fig. 1k).
Cryo-EM shows DICER bound to 5′-G and 5′-U shRNAs
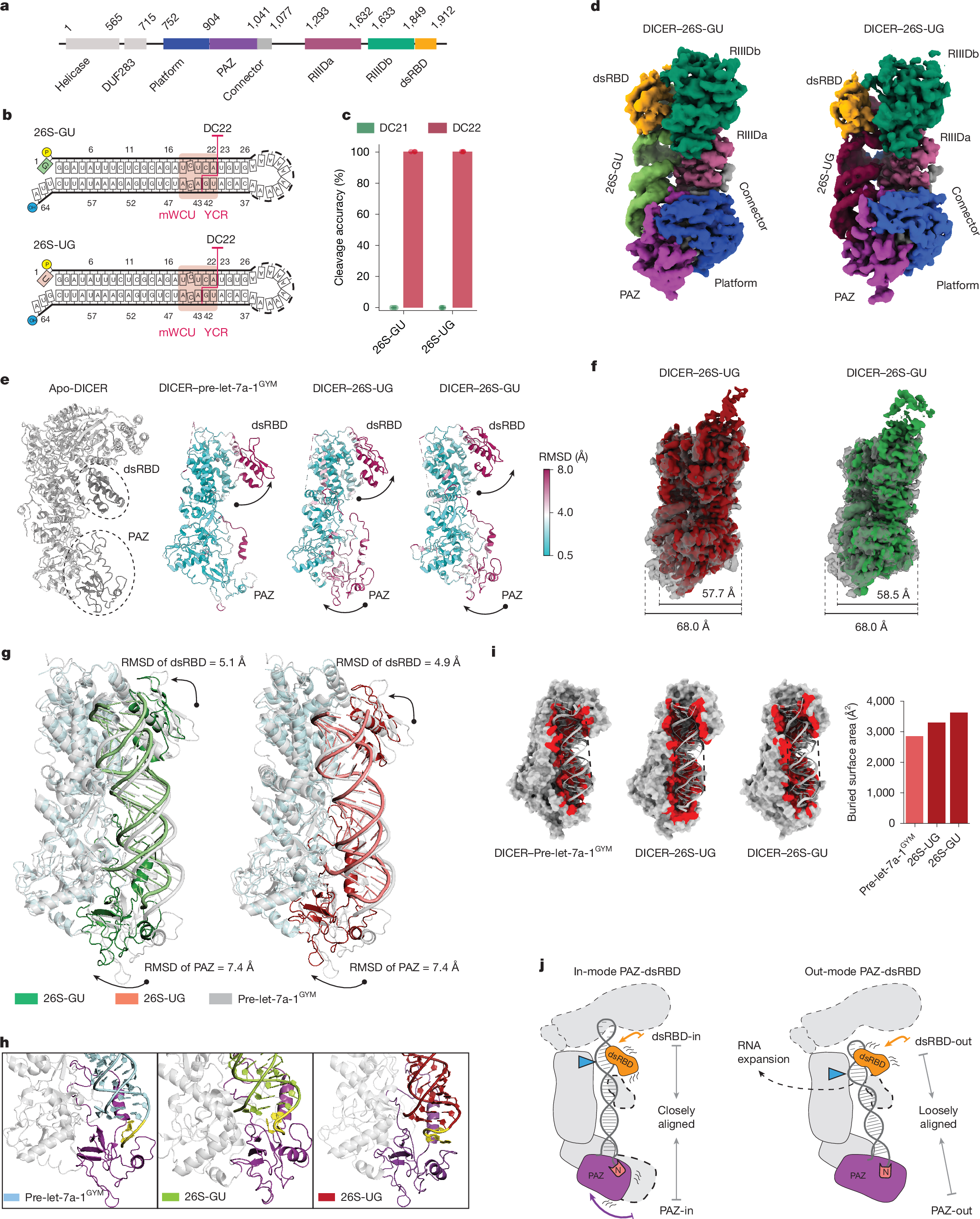
To examine how 5′-G and 5′-U influence DICER cleavage specificity at DC21 and DC22, we solved cryo-EM structures of DICER bound to 5′-G (26S-GU) or 5′-U (26S-UG) shRNAs (Fig. 2a,b). Adding Ca2+ during reconstitution trapped the dicing state by inhibiting catalysis while preserving the cleavage-ready conformation. Cryo-EM maps of the DICER–26S-GU and DICER–26S-UG complexes were obtained at 3.34-Å and 3.37-Å resolution, respectively, as determined by gold-standard Fourier shell correlation (GS-FSC) (Extended Data Fig. 2a–f and Supplementary Table 1). Multiple rounds of two-dimensional (2D) classification selected particles that resembled the dicing-state complex, consistent with previous studies4 (Extended Data Fig. 2a,b). A bias in particle orientation was observed, which is likely to be due to DICER’s asymmetric shape and mass distribution, which might reduce directional resolution and lead to anisotropic reconstructions (Extended Data Fig. 2f). Local-resolution analysis showed slightly higher resolution in protein densities than in RNA densities (Extended Data Fig. 2d). Using the published structural model4 (Protein Data Bank (PDB): 7XW2), we refined atomic structures of DICER in complex with both shRNAs, revealing detailed architecture and domain organization (Extended Data Fig. 2g). The 26S-UG and 26S-GU RNAs included mWCU and YCR motifs, promoting cleavage at DC2235,36, which allowed us to investigate the effects of 5′-nt and RNA motifs on DICER–RNA interactions (Fig. 2c and Extended Data Fig. 3a).
a, Schematic of human DICER domains: DUF283, PAZ, RIIIDa and RIIIDb (RNase III) and dsRBD, with amino acid boundaries labelled. b, Sequences of the 26S-UG and 26S-GU shRNAs that were used in cryo-EM studies. The mWCU and YCR motifs guide DC22 cleavage. c, Cleavage accuracy of DICER for 26S-UG and 26S-GU calculated from three independent assays, showing precise cleavage at DC22. d, Cryo-EM reconstructions of DICER–26S-UG and DICER–26S-GU complexes, with domains colour-coded. e, Structural comparison of DICER complexes reveals conformational shifts in dsRBD and PAZ after RNA binding. RMSD values show significant movement compared with apo-DICER (PDB: 7XW3) and DICER–pre-let-7a-1GYM (PDB: 7XW2). f, Alignment of cryo-EM densities shows that shRNA-bound complexes are more compact (57.7–58.5 Å) than DICER/pre-let-7a-1GYM (68.0 Å). g, PAZ and dsRBD translocate in the DICER–26S-UG and DICER–26S-GU complexes relative to DICER–pre-let-7a-1GYM, with RMSD values of 7.4 Å (PAZ) and around 5.0 Å (dsRBD), aligning the RNA substrate with the catalytic centres. h, PAZ adopts an ‘inner’ conformation in DICER–shRNA complexes, bending the 3′-overhang, unlike the ‘outer’ conformation in DICER–pre-let-7a-1GYM. i, Buried surface area analysis shows increased RNA–DICER interaction in shRNA-bound states, compared with DICER–pre-let-7a-1